library(jsonlite)
# Parse JSON string
response_json <- '{"gauge":"Lees Ferry","flow":12400}'
(data <- fromJSON(response_json))
#> $gauge
#> [1] "Lees Ferry"
#>
#> $flow
#> [1] 12400
# Convert R object back to JSON
data_out <- toJSON(data, pretty = TRUE)
cat(data_out)
#> {
#> "gauge": ["Lees Ferry"],
#> "flow": [12400]
#> }Week 4-2
JSON, GeoJSON, and STAC: Cloud-Native Geospatial Discovery
Learning Objectives
By the end of this week, you will be able to:
- Parse and validate JSON from APIs and cloud storage systems
- Read and write GeoJSON for spatial data exchange in reproducible workflows
- Query STAC catalogs to discover and access petabyte-scale remote sensing datasets
- Download and organize imaging assets directly into analysis pipelines (avoiding manual downloads)
- Apply these tools to real water resource problems (e.g., flood extent mapping, drought monitoring)
Why These Standards Matter for Water Data Science
Modern water resource analysis requires fast, reproducible access to cloud-hosted data:
- JSON: Standard format for REST APIs (USGS, NOAA, NWIS, NASA)
- GeoJSON: Enables spatial data to flow between databases, web services, and R
- STAC: Discovers satellite imagery, model outputs, and time-series data without manual catalog browsing
Real scenario: Analyzing the 2016 Palo, Iowa flood requires Landsat imagery from Sept 24–29, 2016. With STAC + rstac, that’s a 3-line search query. Without it: hours of clicking through NASA/USGS websites.
JSON: The Universal Data Exchange Format
JSON (JavaScript Object Notation) is everywhere in modern data science:
- APIs: USGS NWIS, NOAA, OpenWeather, Mapbox, Google Earth Engine
- Cloud storage: AWS S3 metadata, Azure metadata, Google Cloud Storage
- Databases: NoSQL (MongoDB), time-series (InfluxDB), document stores
- Configuration:
package.json, Docker Compose, Kubernetes
Key advantage: Human-readable + machine-parseable + language-agnostic.
JSON Syntax & Data Types
Objects (key-value pairs):
{
"gauge_id": "09380000",
"name": "Colorado R at Lees Ferry",
"state": "AZ",
"mean_flow_cfs": 12500.5
}Arrays (ordered lists):
Data types: string, number, boolean, null, object, array
Common mistakes: unquoted keys, trailing commas, mismatched quotes
Parsing JSON in R
Use jsonlite::fromJSON() to parse JSON strings or URLs:
Example
jsonlite::fromJSON('https://api.water.usgs.gov/nldi/linked-data/nwissite/USGS-11120000/basin?f=json')
#> $type
#> [1] "FeatureCollection"
#>
#> $features
#> type geometry.type
#> 1 Feature Polygon
#> geometry.coordinates
#> 1 -119.77318, -119.77417, -119.78318, -119.80453, -119.81148, -119.81970, -119.82177, -119.82532, -119.82788, -119.82953, -119.82337, -119.81773, -119.81840, -119.81423, -119.80947, -119.80618, -119.80570, -119.80319, -119.79965, -119.79976, -119.79788, -119.79740, -119.80084, -119.80152, -119.79792, -119.79771, -119.79610, -119.79558, -119.79138, -119.79504, -119.79367, -119.79491, -119.79093, -119.78582, -119.78033, -119.77481, -119.75595, -119.75273, -119.74613, -119.74481, -119.74628, -119.74062, -119.73365, -119.73075, -119.73438, -119.74040, -119.74208, -119.74850, -119.75526, -119.75368, -119.74909, -119.74522, -119.74498, -119.74733, -119.75328, -119.75293, -119.76057, -119.76967, -119.77318, 34.43021, 34.42717, 34.42023, 34.41981, 34.41806, 34.41844, 34.42047, 34.41834, 34.41867, 34.42039, 34.42651, 34.42752, 34.43160, 34.43699, 34.43870, 34.44395, 34.45008, 34.45590, 34.45731, 34.45906, 34.46023, 34.46202, 34.46432, 34.46750, 34.47877, 34.48549, 34.48743, 34.50037, 34.50359, 34.51399, 34.51615, 34.51789, 34.52038, 34.52034, 34.51724, 34.51128, 34.50409, 34.50555, 34.50490, 34.50711, 34.50807, 34.50953, 34.50682, 34.50189, 34.49278, 34.48782, 34.48332, 34.48081, 34.47381, 34.46935, 34.46462, 34.45292, 34.44701, 34.44307, 34.43972, 34.43647, 34.43599, 34.43016, 34.43021GeoJSON: Spatial Data as JSON
GeoJSON encodes geographic features in JSON—standard for web mapping, APIs, and data interchange.
Structure: - Geometry: Points, LineStrings, Polygons (lon/lat coordinates in WGS84) - Properties: Attributes (names, values, measurements) - Feature: Single geometry + properties - FeatureCollection: Multiple features
Point: [-91.793, 42.066] (lon, lat)
Polygon: Ring of coordinates forming closed boundary
GeoJSON Geometry Types
{
"type": "Polygon",
"coordinates": [[
[-91.8, 42.06], [-91.79, 42.07], [-91.78, 42.06], [-91.8, 42.06]
]]
}Key rule: Coordinates are [longitude, latitude] (not lat, lon!)
GeoJSON Feature Example
A single flood extent:
Live Read
basin <- sf::read_sf('https://api.water.usgs.gov/nldi/linked-data/nwissite/USGS-11120000/basin')
basin
#> Simple feature collection with 1 feature and 0 fields
#> Geometry type: POLYGON
#> Dimension: XY
#> Bounding box: xmin: -119.8295 ymin: 34.41806 xmax: -119.7307 ymax: 34.52038
#> Geodetic CRS: WGS 84
#> # A tibble: 1 × 1
#> geometry
#> <POLYGON [°]>
#> 1 ((-119.7732 34.43021, -119.7742 34.42717, -119.7832 34.42023, -119.8045 34.41…
plot(basin)
STAC: Discovering Petabyte-Scale Remote Sensing Data
STAC (SpatioTemporal Asset Catalog) is a standard for describing & discovering imagery:
- Problem: Finding cloud-free Landsat imagery for your study area on a specific date used to require:
- Visiting NASA EarthExplorer or USGS
- Manually filtering by date, cloud cover, location
- Clicking download buttons
- Waiting for large files
- STAC solution: Machine-readable metadata + standard API = programmatic discovery & download
STAC Concepts
- Catalog: Describes a collection of data (e.g., Landsat Collection 2 Level-2)
- Collection: Metadata + spatial/temporal extent for a dataset
- Item: Single asset (one satellite scene), with its own metadata (cloud cover, datetime, bands)
- Asset: Specific file or data product (red band, NIR band, thumbnail, cloud mask)
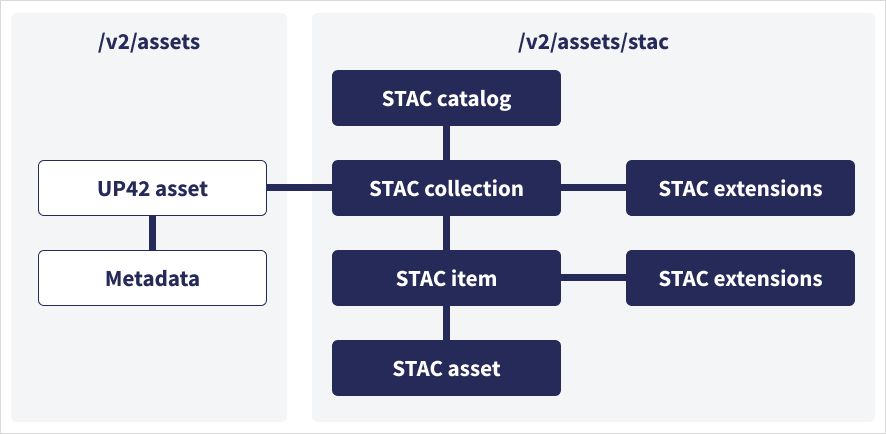
STAC hierarchy: Catalog → Collection → Item → Asset
Key insight: This nested structure lets you query at any level (find catalogs, browse collections, search for items, download specific assets).
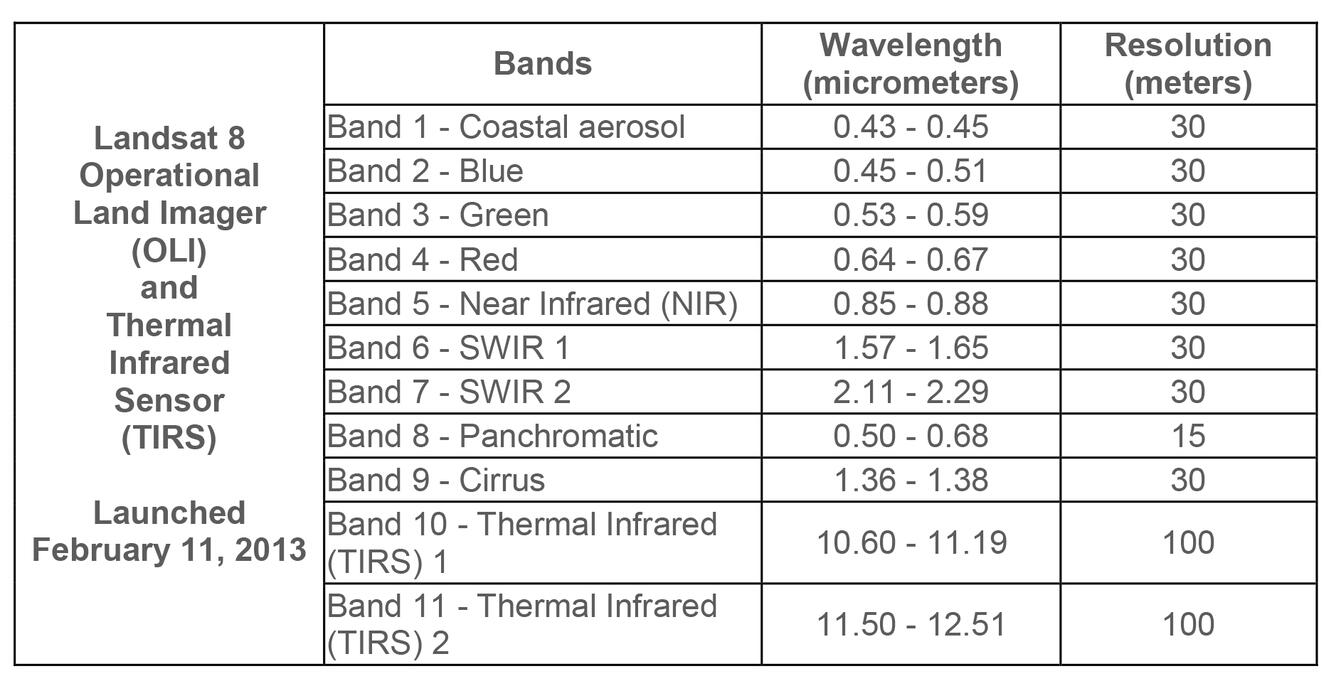
STAC Item Anatomy (Landsat Example)
{
"type": "Feature",
"id": "LC08_L2SP_025031_20160926_02_T1",
"stac_version": "1.0.0",
"geometry": { "type": "Polygon", "coordinates": [...] },
"properties": {
"datetime": "2016-09-26T16:47:00Z",
"platform": "landsat-8",
"instruments": ["OLI", "TIRS"],
"eo:cloud_cover": 2.1
},
"assets": {
"red": { "href": "s3://...LC08_red.TIF" },
"nir": { "href": "s3://...LC08_nir.TIF" }
}
}Key insight: Assets are URLs pointing to cloud storage. A STAC item is essentially metadata + pointers to files.
Catalog
Querying STAC with rstac: Palo 2016 Flood
The Palo flood occurred Sept 26, 2016. Search for Landsat imagery covering that event:
Collections
(
search <- stac(stac_ep) |>
stac_search(
collections = "landsat-c2-l2",
)|>
get_request()
)
#> ###Items
#> - features (250 item(s)):
#> - LC09_L2SP_090091_20260414_02_T2
#> - LC09_L2SP_090090_20260414_02_T1
#> - LC09_L2SP_090089_20260414_02_T1
#> - LC09_L2SP_090088_20260414_02_T1
#> - LC09_L2SP_090087_20260414_02_T1
#> - LC09_L2SP_090086_20260414_02_T1
#> - LC09_L2SP_090085_20260414_02_T1
#> - LC09_L2SP_090084_20260414_02_T1
#> - LC09_L2SP_090083_20260414_02_T1
#> - LC09_L2SP_090082_20260414_02_T1
#> - ... with 240 more feature(s).
#> - assets:
#> ang, atran, blue, cdist, coastal, drad, emis, emsd, green, lwir11, mtl.json, mtl.txt, mtl.xml, nir08, qa, qa_aerosol, qa_pixel, qa_radsat, red, rendered_preview, swir16, swir22, tilejson, trad, urad
#> - item's fields:
#> assets, bbox, collection, geometry, id, links, properties, stac_extensions, stac_version, typeSpatial Filter
(
search <- stac(stac_ep) |>
stac_search(
collections = "landsat-c2-l2",
bbox = st_bbox(bbox_palo),
limit = 10
)|>
get_request()
)
#> ###Items
#> - features (10 item(s)):
#> - LC09_L2SP_025031_20260407_02_T1
#> - LC08_L2SP_025031_20260330_02_T1
#> - LC09_L2SP_026031_20260329_02_T1
#> - LC09_L2SP_025031_20260322_02_T1
#> - LC08_L2SP_026031_20260321_02_T1
#> - LC08_L2SP_025031_20260314_02_T1
#> - LC09_L2SP_026031_20260313_02_T1
#> - LC09_L2SP_025031_20260306_02_T1
#> - LC08_L2SP_026031_20260305_02_T2
#> - LC08_L2SP_025031_20260226_02_T1
#> - assets:
#> ang, atran, blue, cdist, coastal, drad, emis, emsd, green, lwir11, mtl.json, mtl.txt, mtl.xml, nir08, qa, qa_aerosol, qa_pixel, qa_radsat, red, rendered_preview, swir16, swir22, tilejson, trad, urad
#> - item's fields:
#> assets, bbox, collection, geometry, id, links, properties, stac_extensions, stac_version, typeTemporal Filter
(
search <- stac(stac_ep) |>
stac_search(
collections = "landsat-c2-l2",
bbox = st_bbox(bbox_palo),
datetime = "2016-09-24/2016-09-29",
limit = 10
)|>
get_request()
)
#> ###Items
#> - features (2 item(s)):
#> - LC08_L2SP_025031_20160926_02_T1
#> - LE07_L2SP_026031_20160925_02_T1
#> - assets:
#> ang, atmos_opacity, atran, blue, cdist, cloud_qa, coastal, drad, emis, emsd, green, lwir, lwir11, mtl.json, mtl.txt, mtl.xml, nir08, qa, qa_aerosol, qa_pixel, qa_radsat, red, rendered_preview, swir16, swir22, tilejson, trad, urad
#> - item's fields:
#> assets, bbox, collection, geometry, id, links, properties, stac_extensions, stac_version, typeGet Request
The GET request only returns the JSON catalog showing were data is located this is “free” to all.
To actually get the assests from the URL, some providers require a key, some an account, and some just an authentication
Microsoft is an example of authenticated but free to use!
Signed URLs & Accessing Planetary Computer Data
Microsoft Planetary Computer hosts large datasets (Landsat, Sentinel, Copernicus, MODIS, etc.) and requires signed URLs for access. The rstac package handles this automatically:
# Sign all asset URLs for the first item
item_signed <- search |>
items_sign(rstac::sign_planetary_computer())
item_signed$features[[1]]$assets$green$href # Now this URL is signed and ready for download
#> [1] "https://landsateuwest.blob.core.windows.net/landsat-c2/level-2/standard/oli-tirs/2016/025/031/LC08_L2SP_025031_20160926_20200906_02_T1/LC08_L2SP_025031_20160926_20200906_02_T1_SR_B3.TIF?st=2026-04-14T21%3A20%3A17Z&se=2026-04-15T22%3A05%3A17Z&sp=rl&sv=2025-07-05&sr=c&skoid=9c8ff44a-6a2c-4dfb-b298-1c9212f64d9a&sktid=72f988bf-86f1-41af-91ab-2d7cd011db47&skt=2026-04-15T00%3A02%3A34Z&ske=2026-04-22T00%3A02%3A34Z&sks=b&skv=2025-07-05&sig=%2BOTGhGiKwIXrnO6kU4uaGmg5z1A9XFdY001GaFccOYs%3D"This avoids keeping credentials in your code—much more secure!
Safe Download: Check File Size First
Before downloading large band files (100s of MB), inspect the file size with a HEAD request:
# Example: safe download pattern
file_url <- item_signed[[1]]$assets$green$href
# HEAD request: get metadata without downloading
h <- httr::HEAD(file_url)
if (httr::status_code(h) == 200) {
file_size_mb <- as.numeric(httr::headers(h)$`content-length`) / 1e6
cat("File size:", file_size_mb, "MB\n")
# Only download if < 500 MB
if (file_size_mb < 500) {
httr::GET(file_url, httr::write_disk("band_red.TIF", overwrite = TRUE))
cat("Downloaded successfully\n")
} else {
cat("File too large; consider downloading selectively\n")
}
}Direct read
vsi <- terra::rast(item_signed$features[[1]]$assets$green$href)
vsi
#> class : SpatRaster
#> size : 7801, 7681, 1 (nrow, ncol, nlyr)
#> resolution : 30, 30 (x, y)
#> extent : 518085, 748515, 4506885, 4740915 (xmin, xmax, ymin, ymax)
#> coord. ref. : WGS 84 / UTM zone 15N (EPSG:32615)
#> source : LC08_L2SP_025031_20160926_20200906_02_T1_SR_B3.TIF
#> name : LC08_L2SP_025031_20160926_20200906_02_T1_SR_B3Assisted Asset Download
list.files( path = "../labs/data",
pattern = "\\.TIF$",
recursive = TRUE,
full.names = TRUE)
#> [1] "../labs/data/landsat-c2/level-2/standard/oli-tirs/2016/025/031/LC08_L2SP_025031_20160926_20200906_02_T1/LC08_L2SP_025031_20160926_20200906_02_T1_SR_B1.TIF"
#> [2] "../labs/data/landsat-c2/level-2/standard/oli-tirs/2016/025/031/LC08_L2SP_025031_20160926_20200906_02_T1/LC08_L2SP_025031_20160926_20200906_02_T1_SR_B2.TIF"
#> [3] "../labs/data/landsat-c2/level-2/standard/oli-tirs/2016/025/031/LC08_L2SP_025031_20160926_20200906_02_T1/LC08_L2SP_025031_20160926_20200906_02_T1_SR_B3.TIF"
#> [4] "../labs/data/landsat-c2/level-2/standard/oli-tirs/2016/025/031/LC08_L2SP_025031_20160926_20200906_02_T1/LC08_L2SP_025031_20160926_20200906_02_T1_SR_B4.TIF"
#> [5] "../labs/data/landsat-c2/level-2/standard/oli-tirs/2016/025/031/LC08_L2SP_025031_20160926_20200906_02_T1/LC08_L2SP_025031_20160926_20200906_02_T1_SR_B5.TIF"
#> [6] "../labs/data/landsat-c2/level-2/standard/oli-tirs/2016/025/031/LC08_L2SP_025031_20160926_20200906_02_T1/LC08_L2SP_025031_20160926_20200906_02_T1_SR_B6.TIF"From STAC to Analysis: The Full Workflow
Query STAC Inspect Sign URLs Download Load & Analyze
Find scenes → Metadata/CC → Authenticate → Selectively → NDVI, floods, etc.
↓ ↓ ↓ ↓ ↓
bbox, date cloud cover secured tokens 30m TIFFs terra raster
collection available valid to disk ensemble
bands ~15 min only needed mappingLab 4 & this walkthrough execute all five steps for Palo 2016 flood.
- Query STAC → find scenes matching criteria (date, location, cloud cover)
- Inspect metadata → confirm cloud cover, available bands, data quality
- Sign URLs → authenticate access to cloud-hosted files
- Download selectively → you often need only 1–2 bands, not entire scenes
- Load into
terra→ treat as raster for analysis (NDVI, flood mapping, etc.)
Why This Matters for Water Science
Pre-STAC workflow (still common):
Click directories on EarthExplorer
Download entire scenes (10+ GB each)
Manual metadata tracking
Months to assemble regional datasets
With STAC + rstac:
Query programmatically in seconds
Download only what you need (individual bands, 50–100 MB)
Metadata is structured, version-controlled
Automated multi-year, multi-sensor analyses feasible
Real-World STAC Catalogs for Water
- Microsoft Planetary Computer: 50+ datasets (Landsat, Sentinel-1/2, MODIS, ASTER, Climate models)
- AWS Registry of Open Data: Open-access Landsat, Sentinel, NOAA, others
- Google Earth Engine: Similar concept (not STAC-native, but queryable)
- USGS OpenData: Some products via STAC
- CBERS/INPE (Brazil): Open satellite data via STAC
All queryable programmatically from R with rstac.
Key Takeaways
- JSON is the lingua franca of modern APIs—parsing it in R is essential
- GeoJSON enables you to encode, share, and analyze spatial data portably
- STAC unlocks programmatic access to massive remote sensing archives without manual downloads
- Together, these unlock automated multi-year, multi-region analyses
- For water science: combine satellite data with ground-truth (gauges, models) to answer critical operational questions
Complete Walkthrough:
Palo 2016 Flood Analysis
Step 1: Define AOI and Temporal Window
Complete Walkthrough: STAC Query (with signing)
# Connect to Microsoft Planetary Computer
stac_query <- stac("https://planetarycomputer.microsoft.com/api/stac/v1") %>%
stac_search(
collections = "landsat-c2-l2",
datetime = temporal_range,
bbox = st_bbox(palo),
limit = 1
) %>%
get_request() %>%
items_sign(sign_planetary_computer())
stac_query
#> ###Items
#> - features (1 item(s)):
#> - LC08_L2SP_025031_20160926_02_T1
#> - assets:
#> ang, atran, blue, cdist, coastal, drad, emis, emsd, green, lwir11, mtl.json, mtl.txt, mtl.xml, nir08, qa, qa_aerosol, qa_pixel, qa_radsat, red, rendered_preview, swir16, swir22, tilejson, trad, urad
#> - item's fields:
#> assets, bbox, collection, geometry, id, links, properties, stac_extensions, stac_version, type
# Inspect the result
cat("Item ID:", stac_query$features[[1]]$id, "\n")
#> Item ID: LC08_L2SP_025031_20160926_02_T1
cat("Date:", stac_query$features[[1]]$properties$datetime, "\n")
#> Date: 2016-09-26T16:47:41.390781Z
cat("Cloud cover:", stac_query$features[[1]]$properties$`eo:cloud_cover`, "%\n")
#> Cloud cover: 0.01 %Complete Walkthrough: Load Pre-Downloaded Data
Since data is already in labs/data, load directly from the local directory:
# List downloaded Landsat band files (TIF format)
raster_files <- list.files(
path = "../labs/data",
pattern = "*.TIF$",
full.names = TRUE,
recursive = TRUE
)
# Sort to ensure consistent band order
raster_files <- sort(raster_files)[1:6]
# Stack into a single SpatRaster object
landsat_stack <- rast(raster_files)
band_names <- c('coastal', 'blue', 'green', 'red', 'nir08', 'swir16')
landsat_stack <- setNames(landsat_stack, band_names)
landsat_stack
#> class : SpatRaster
#> size : 7801, 7681, 6 (nrow, ncol, nlyr)
#> resolution : 30, 30 (x, y)
#> extent : 518085, 748515, 4506885, 4740915 (xmin, xmax, ymin, ymax)
#> coord. ref. : WGS 84 / UTM zone 15N (EPSG:32615)
#> sources : LC08_L2SP_025031_20160926_20200906_02_T1_SR_B1.TIF
#> LC08_L2SP_025031_20160926_20200906_02_T1_SR_B2.TIF
#> LC08_L2SP_025031_20160926_20200906_02_T1_SR_B3.TIF
#> ... and 3 more sources
#> names : coastal, blue, green, red, nir08, swir16Complete Walkthrough: Crop to AOI
# Transform AOI to match raster CRS (UTM zone 15N)
palo_utm <- bbox_palo %>%
st_transform(crs(landsat_stack))
# Crop the full scene to study area (saves memory, speeds processing)
landsat_crop <- crop(landsat_stack, palo_utm)
cat("\nCropped extent:\n")
#>
#> Cropped extent:
print(ext(landsat_crop))
#> SpatExtent : 598185, 601065, 4655805, 4659615 (xmin, xmax, ymin, ymax)
cat("Cropped dimensions:", dim(landsat_crop), "\n")
#> Cropped dimensions: 127 96 6Complete Walkthrough: Visualize Band Combinations
Natural Color (RGB) — What the Human Eye Would See
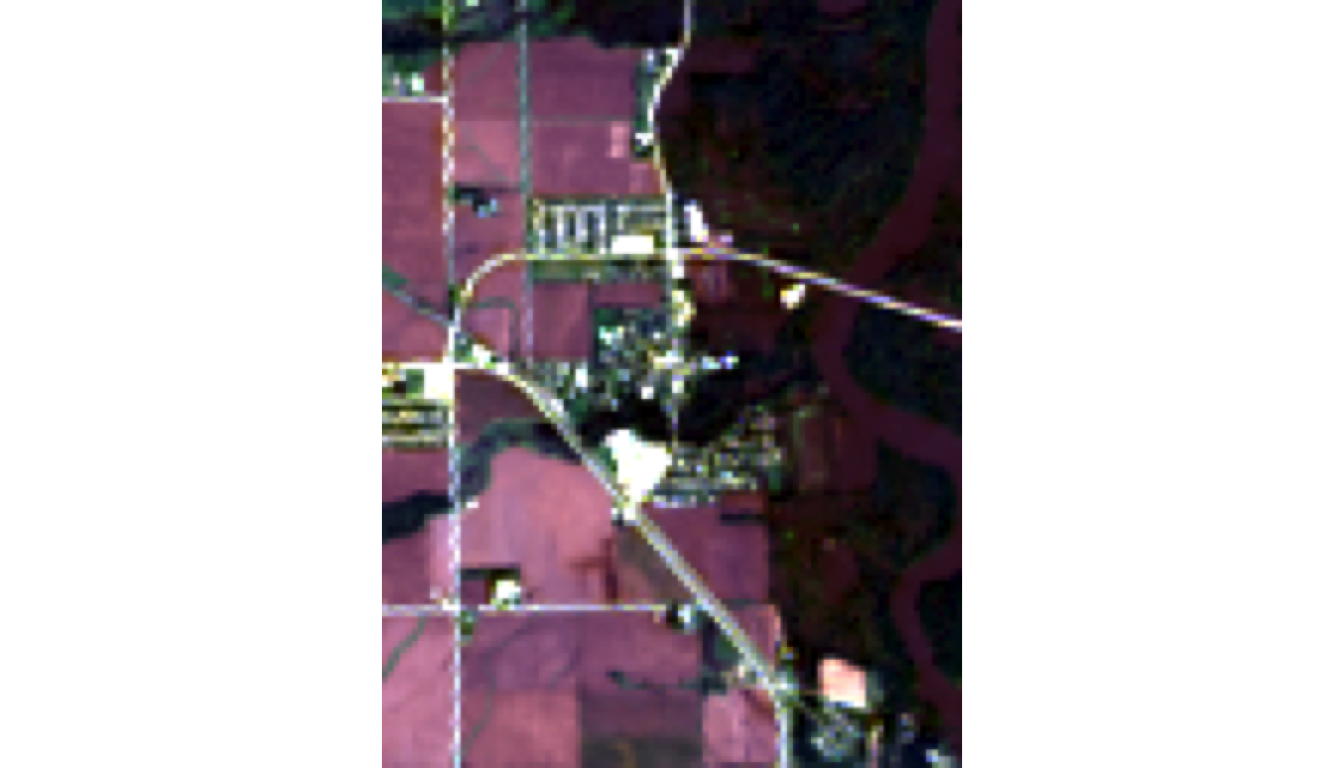

Visible: Water (dark), vegetation (green), urban (gray/tan), clouds (white)
Limitation: Flooding can look similar to vegetation in RGB alone.
Visualize: Color Infrared (Vegetation)
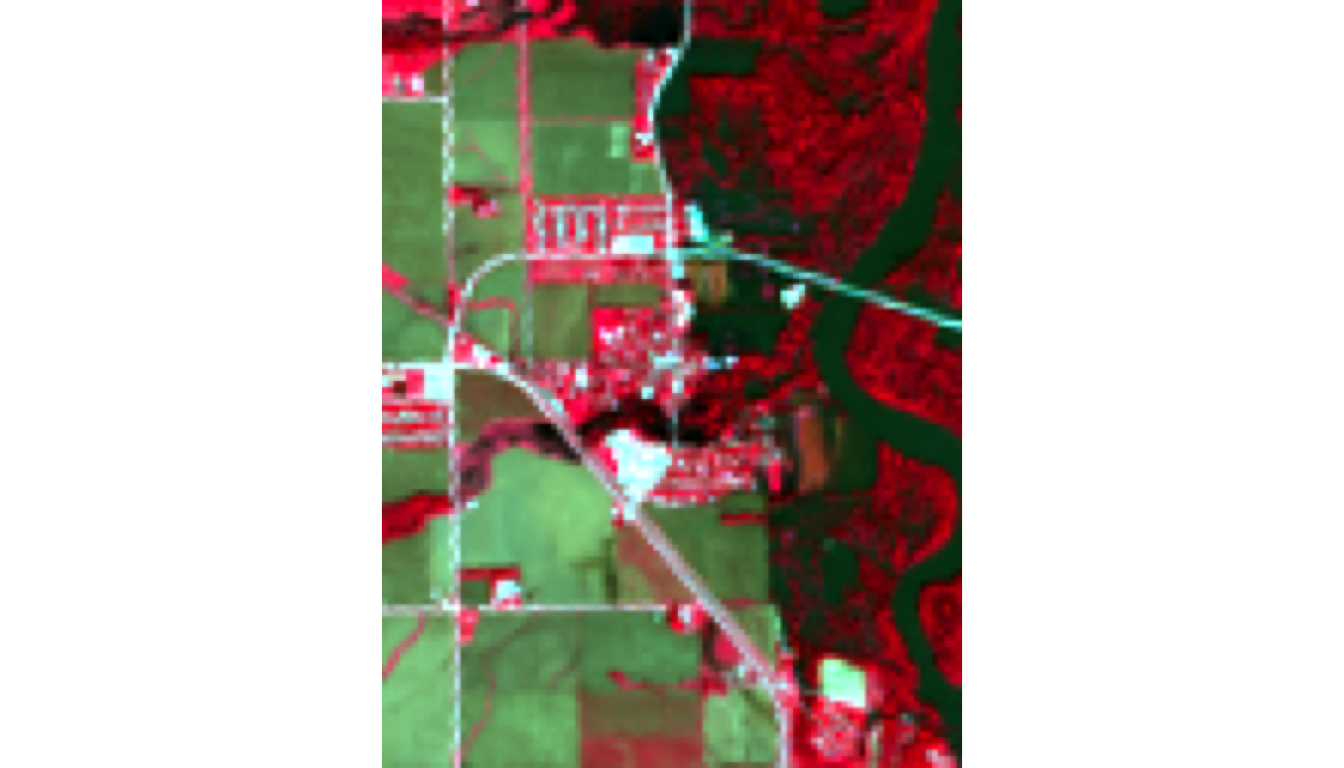

Visible: - Bright red/magenta = healthy vegetation (strong NIR reflectance) - Dark blue/black = water - Gray/tan = soil/urban
Why NIR matters: Vegetation reflects ~50% of NIR light; water absorbs it. This makes forests “glow” red.
Visualize: Water-Focused False Color
plotRGB(landsat_crop, r = 5, g = 6, b = 4, stretch = "lin",
main = "Water Focus: NIR-SWIR1-Red\n(Dark blue = open water/flooding)")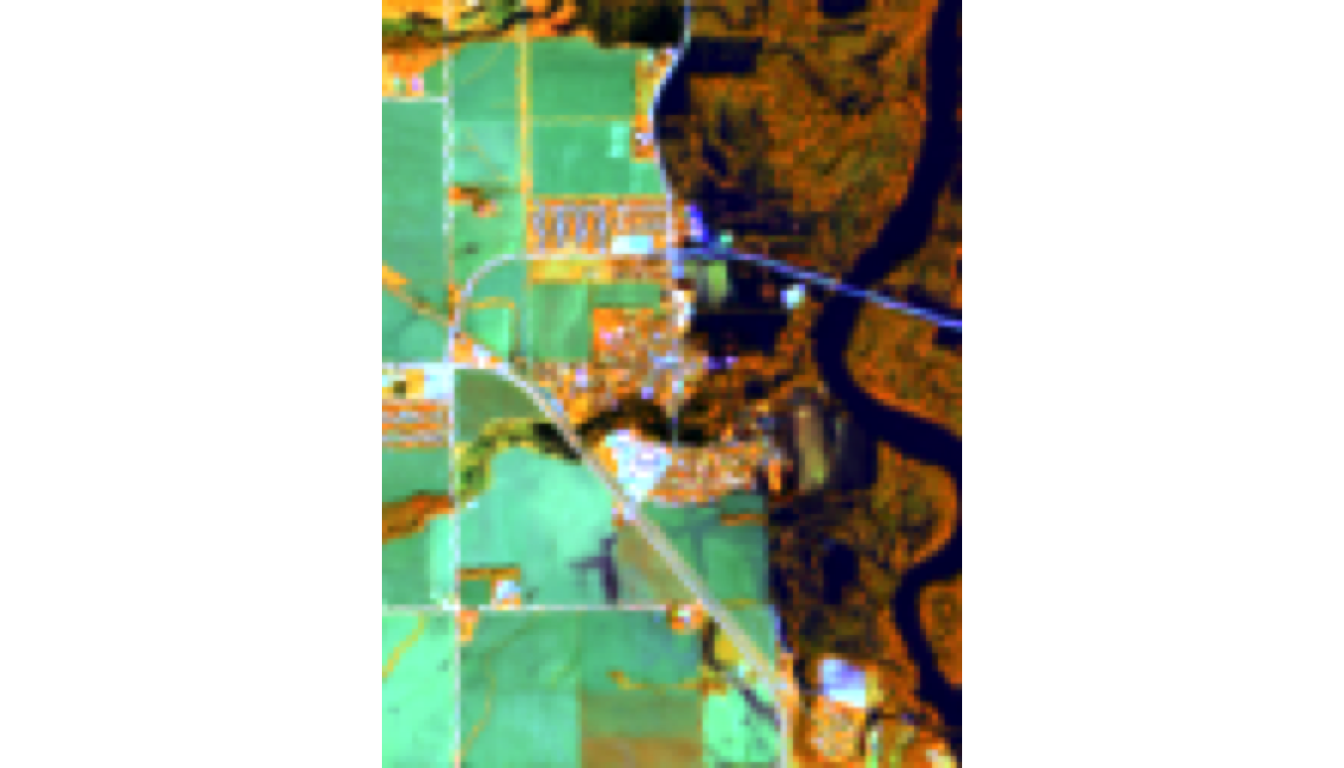
Visible: - Dark blue/black = open water, flooded areas (absorbs NIR + SWIR) - Bright colors = vegetation & land - Often shows flooding more clearly than natural color
Why SWIR1 matters: Water absorbs SWIR; vegetation reflects it. This combo best delineates water boundaries.

Complete Walkthrough: NDVI
$ NDVI = $
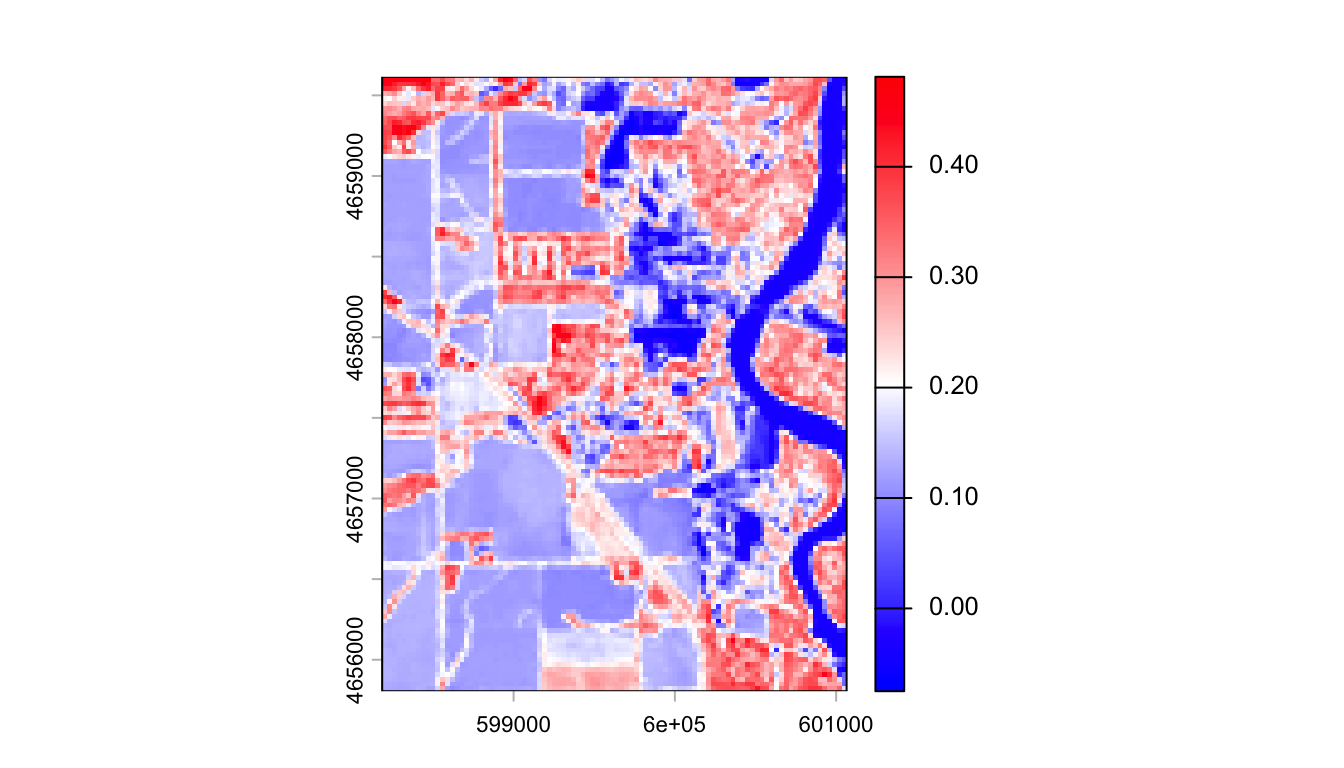
Thresholding to Binary Flood Maps
NDVI < 0 typically indicates water. We can apply this threshold to create a binary flood map:
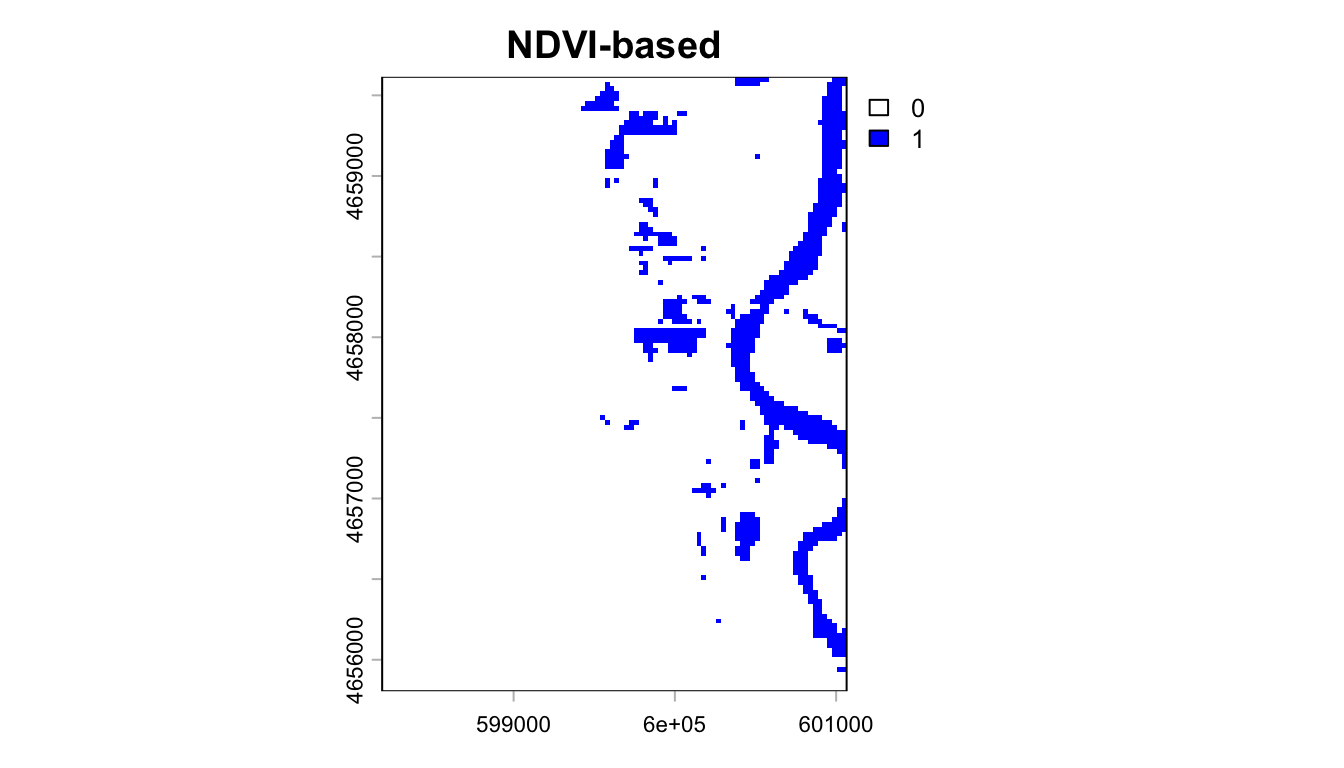
Imapct Assessment:
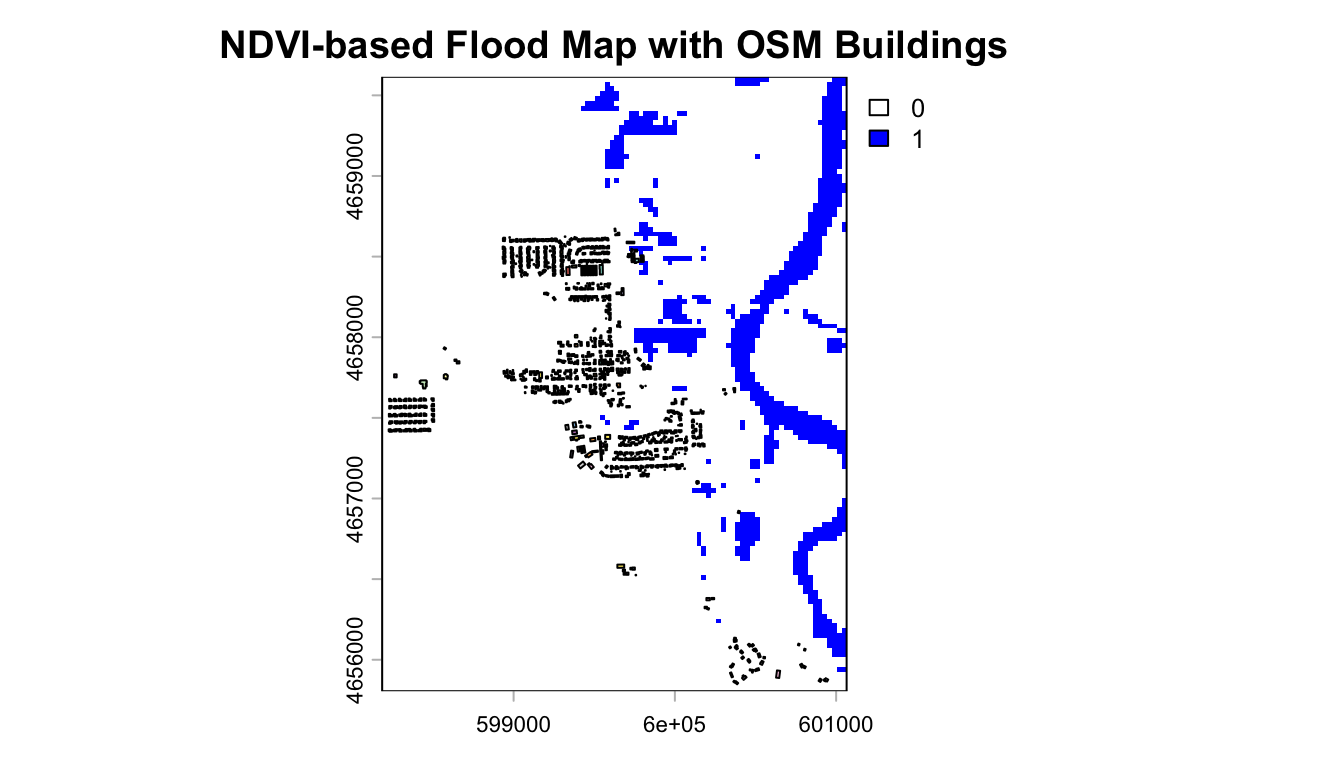
Extract maximum NDVI flood value for each building:
Key Insights from the Walkthrough
- STAC query identified the exact cloud-free Landsat 8 scene (Sept 26, 2016)
- Local files (pre-downloaded in
labs/data) eliminated long download wait - Band stacking combined 6 spectral bands into a single raster object
- Color composites revealed features invisible in natural color (vegetation, water)
- Indices + thresholding converted continuous values to binary flood/no-flood
- Ensemble mapping quantified confidence by counting method agreement
Result: A rigorous, reproducible flood map derived from open satellite data, validated against known drone footage.
Further Reading & Resources
- JSON Spec: https://www.json.org/
- GeoJSON RFC 7946: https://tools.ietf.org/html/rfc7946
- STAC Specification: https://stacspec.org/
- Planetary Computer STAC API: https://planetarycomputer.microsoft.com/api/stac/v1
rstacPackage: https://cran.r-project.org/package=rstac
