Week 5
Machine Learning Part 1
Modeling (Machine Learning)

By the end of today you will be able to…
- Summarize a dataset’s structure and distributions using EDA tools (
skimr,ggpubr,visdat) - Assess whether data meets normality assumptions and choose an appropriate response (transform, nonparametric, bootstrap)
- Explain the role of training, testing, and resampling splits in a modeling workflow
- Specify a model with
parsnipby choosing a type, engine, and mode - Fit a model inside a
workflowand evaluate it withyardstickmetrics - Compare multiple models at once with
workflow_setsover cross-validation folds
Today’s running example: “Can we predict whether a 6,000-acre hexagon in Washington is forested?”
What is machine learning?
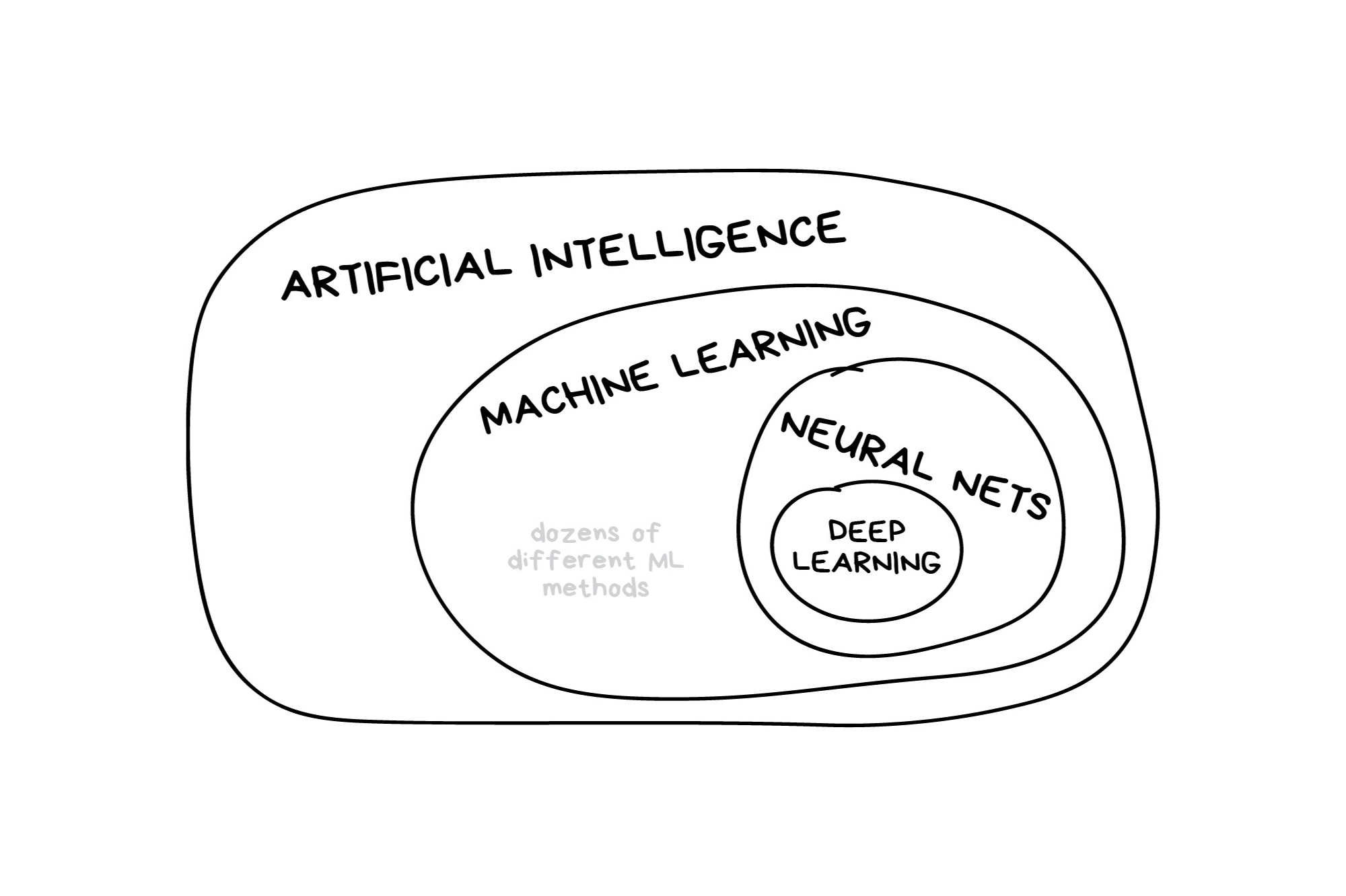
Illustration credit: https://vas3k.com/blog/machine_learning/
What is machine learning? (2025 edition)
Note
In the early 2010s, “Artificial intelligence” (AI) was largely synonymous with what we’ll refer to as “machine learning” in this workshop. In the late 2010s and early 2020s, AI usually referred to deep learning methods. Since the release of ChatGPT in late 2022, “AI” has come to also encompass large language models (LLMs) / generative models.
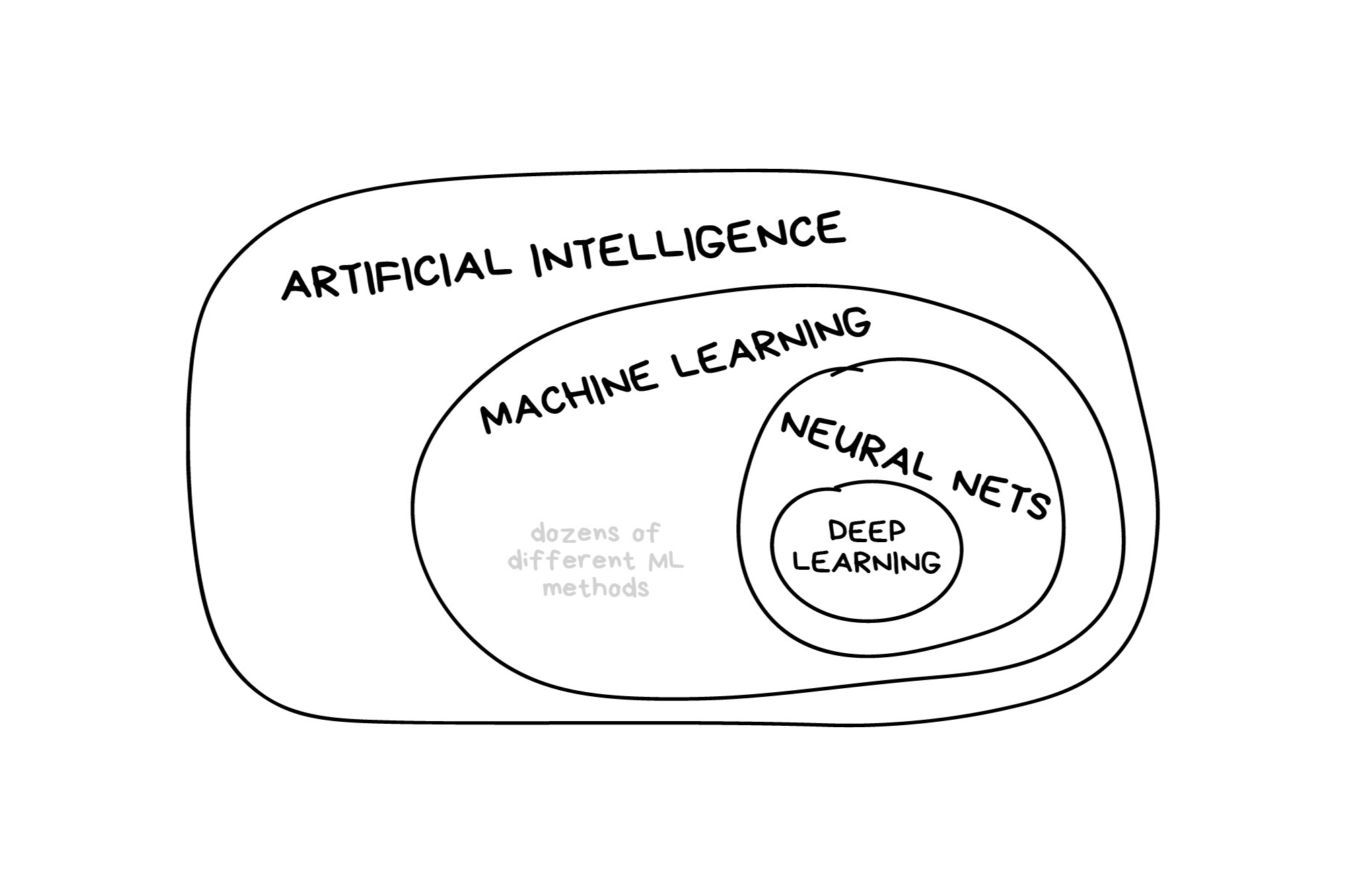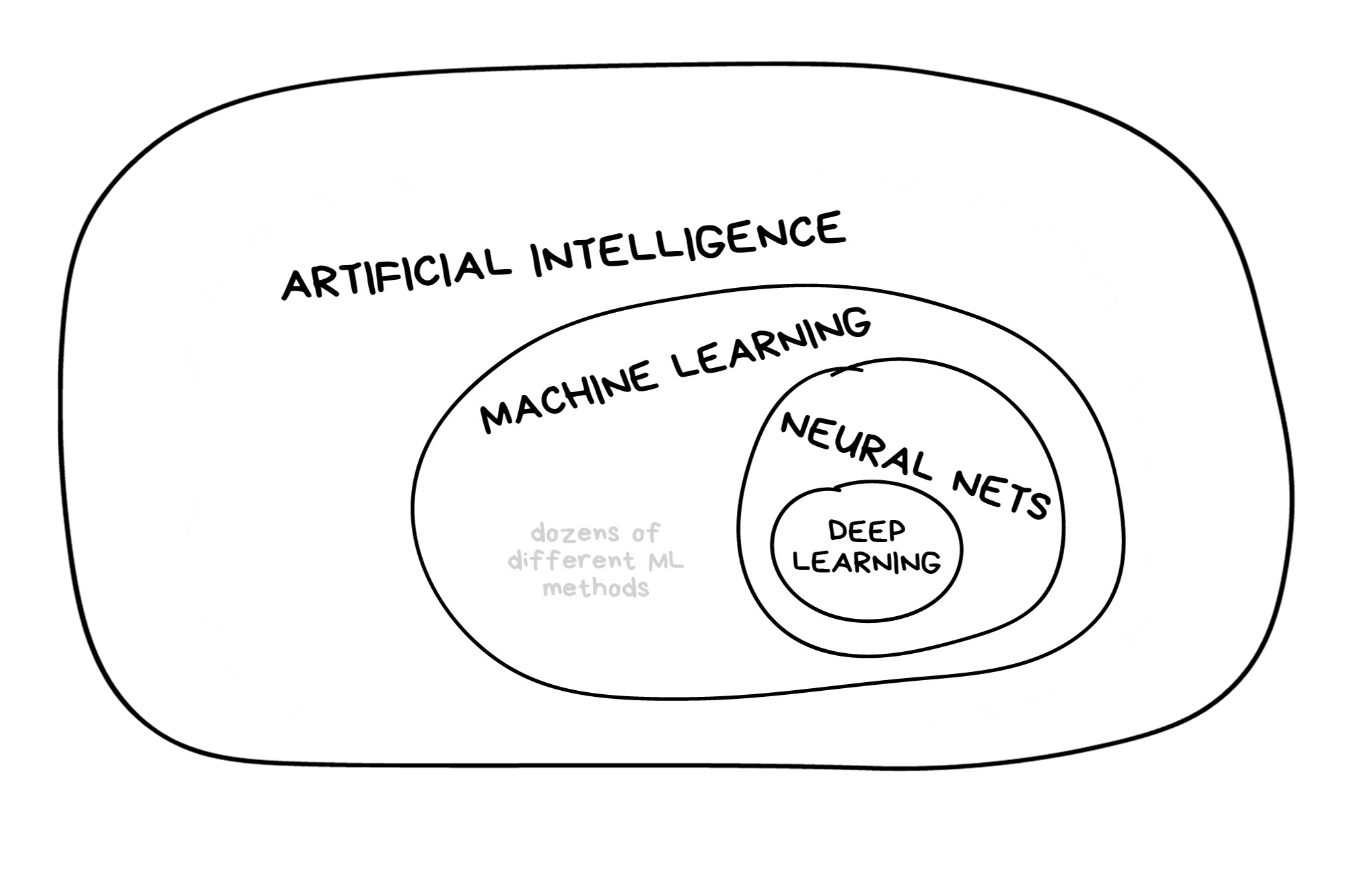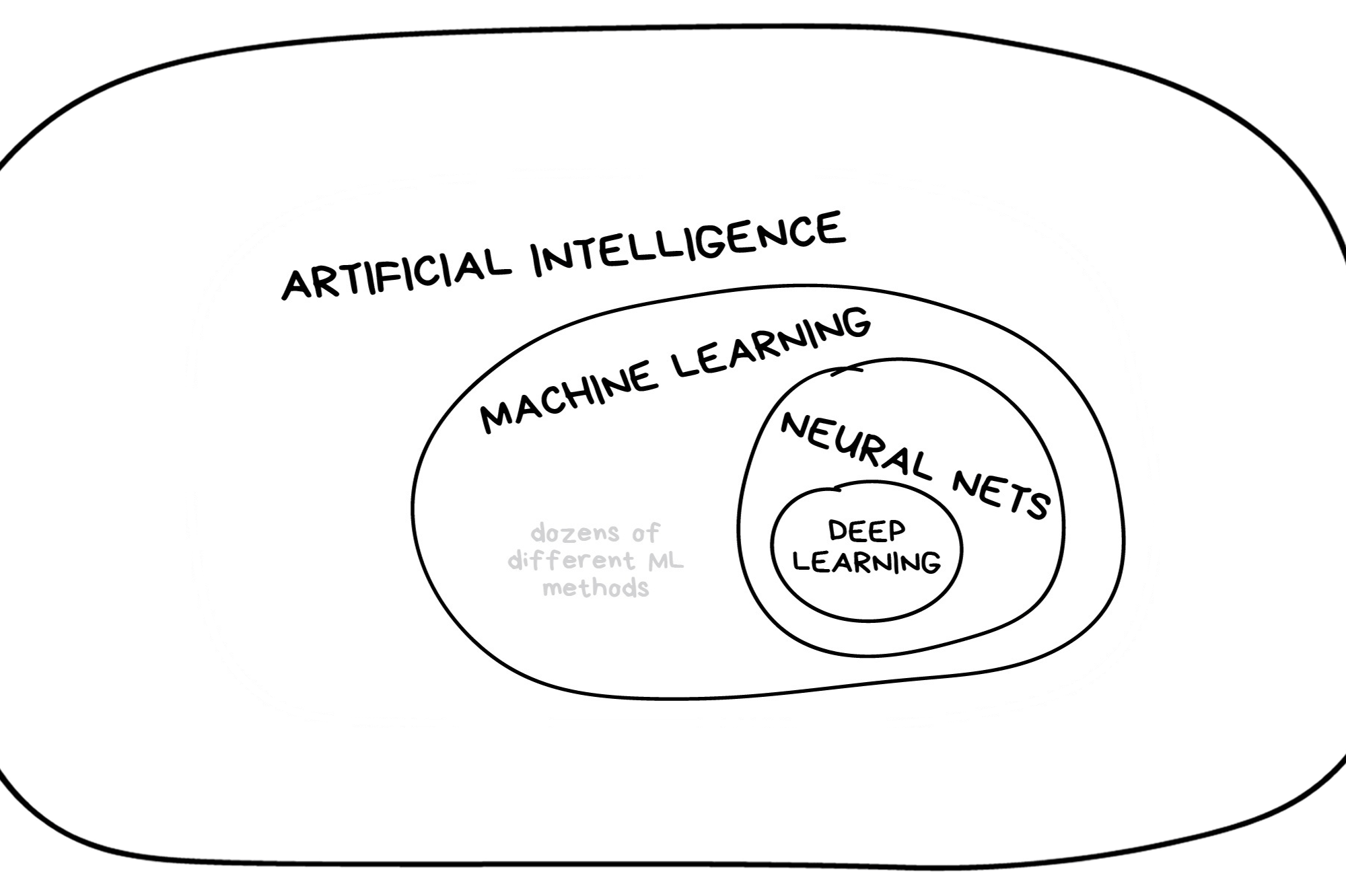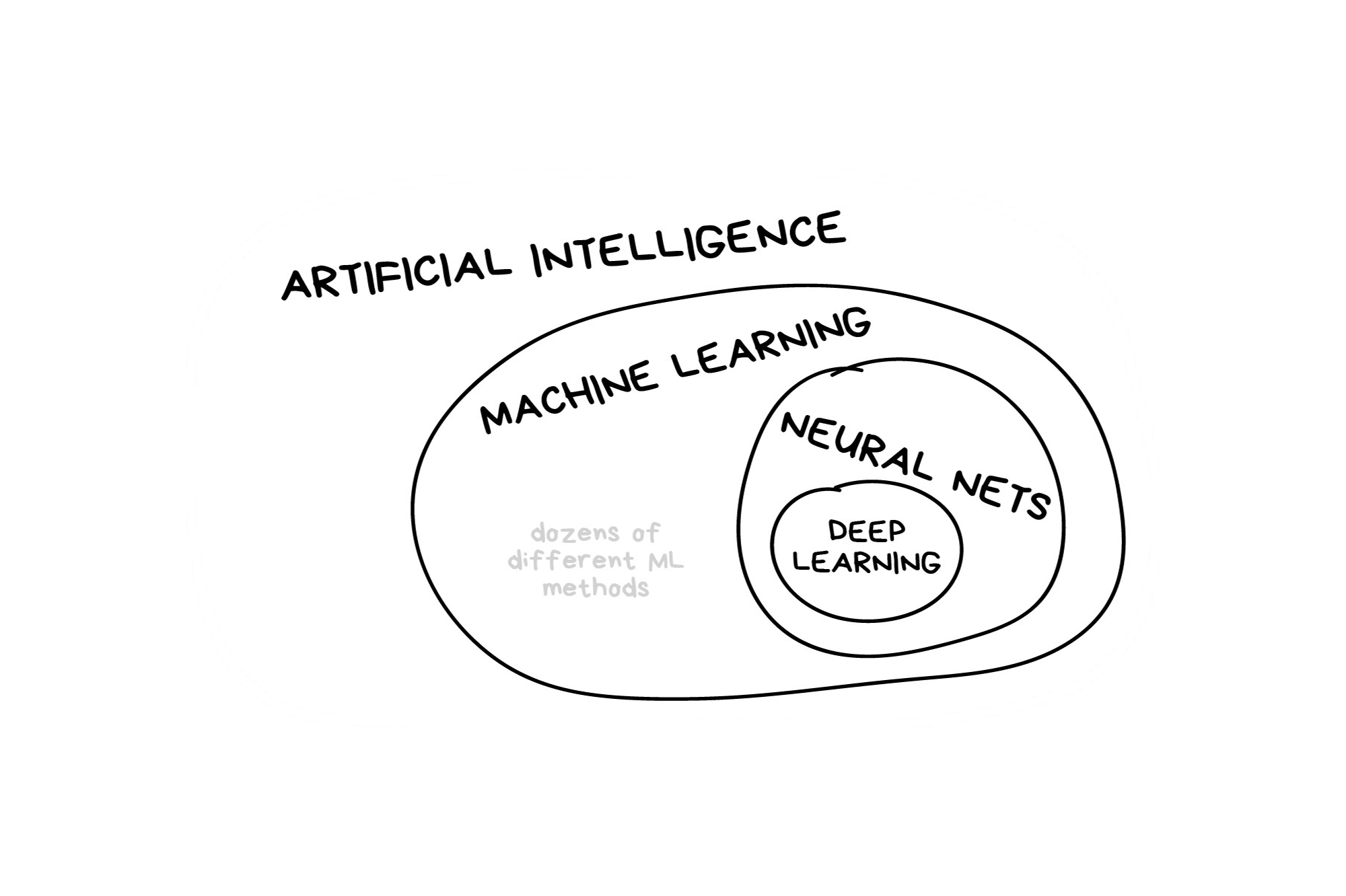
Illustration credit: https://vas3k.com/blog/machine_learning/
Classic Conceptual Model
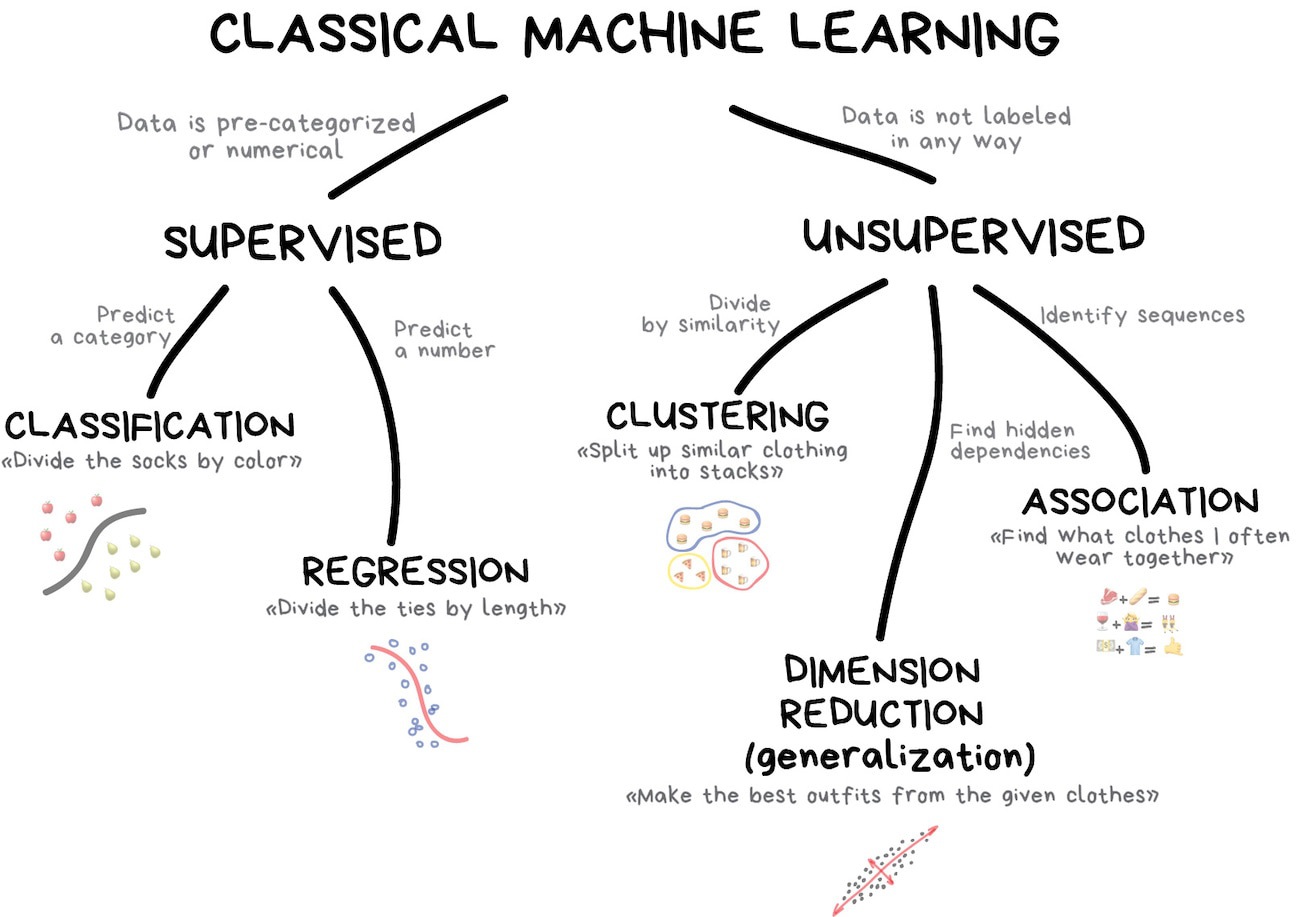
Illustration credit: https://vas3k.com/blog/machine_learning/
The big picture: Road map
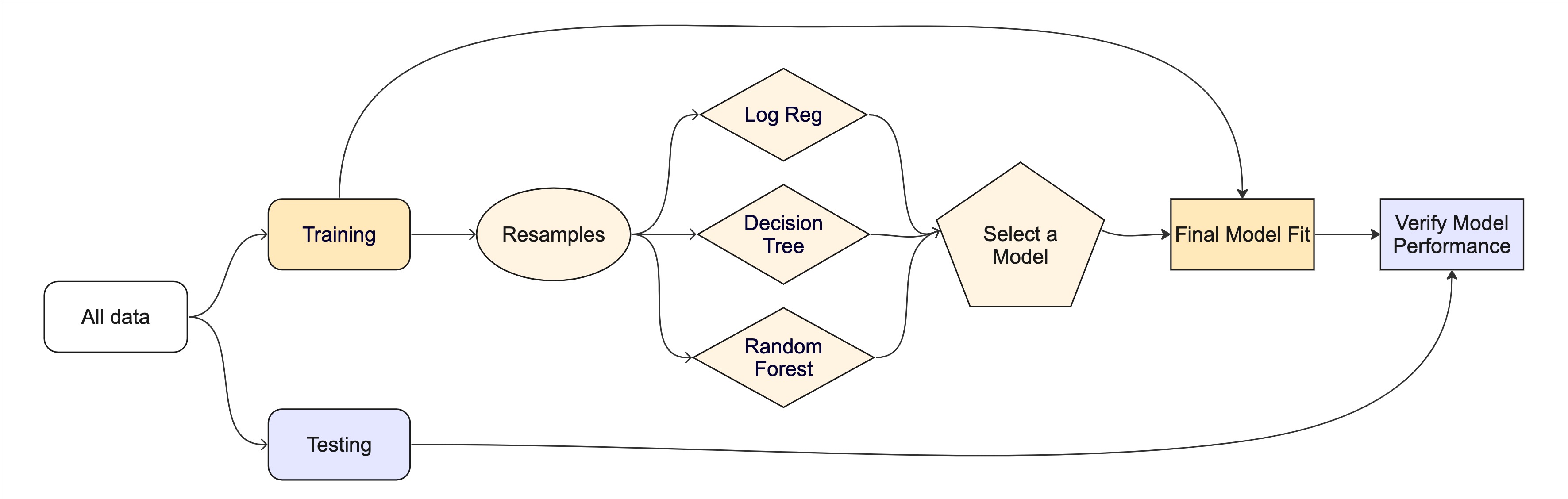
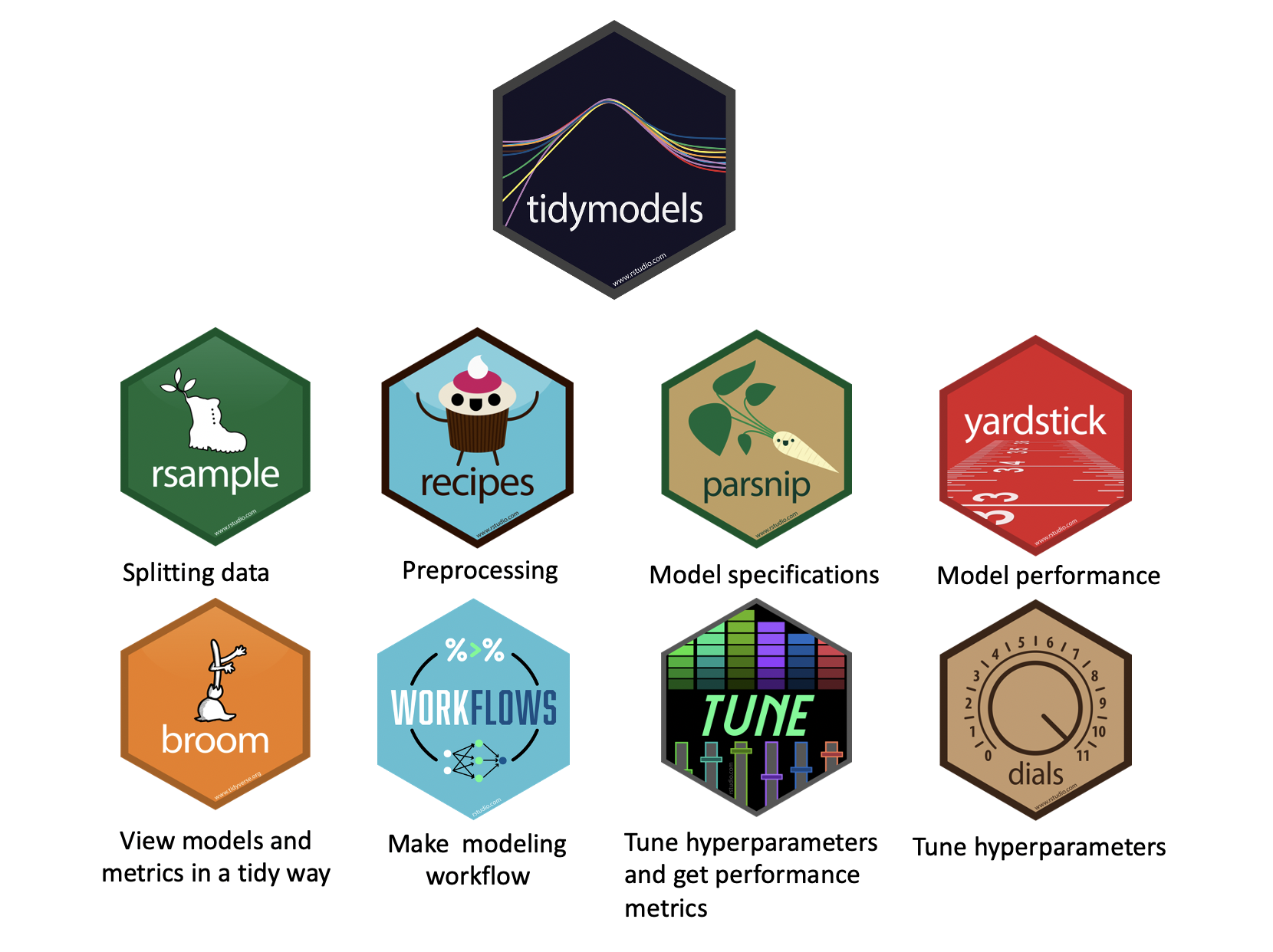
What is tidymodels?
- An R package ecosystem for modeling and machine learning
- Built on the
tidyverseprinciples (consistent syntax, modular design) - Provides a structured workflow for preprocessing, model fitting, and evaluation
- Includes packages for model specification, preprocessing, resampling, tuning, and evaluation
- Promotes best practices for reproducibility and efficiency
- Offers a unified interface for different models and tasks
Key tidymodels Packages
📦 recipes: Feature engineering and preprocessing
📦 parsnip: Unified interface for model specification
📦 workflows: Streamlined modeling pipelines
📦 tune: Hyperparameter tuning
📦 rsample: Resampling and validation
📦 yardstick: Model evaluation metrics
Typical Workflow in tidymodels
1️⃣ Load Package & Data
2️⃣ Preprocess Data (recipes)
3️⃣ Define Model (parsnip)
4️⃣ Create a Workflow (workflows)
5️⃣ Train & Tune (tune)
6️⃣ Evaluate Performance (yardstick)
Recap: lm() in R
You’ve used lm() before — this slide frames it as the foundation for everything broom and tidymodels build on top of.
lm() fits a linear model by ordinary least squares.
Key arguments:
formula—response ~ predictors(.= all columns)data— the training data frame
Common follow-up functions:
| Function | What it returns |
|---|---|
summary(model) |
Coefficients, R², F-statistic |
coef(model) |
Named coefficient vector |
fitted(model) |
In-sample predictions |
residuals(model) |
Raw residuals |
mod <- lm(body_mass_g ~ bill_length_mm + species,
data = drop_na(penguins))
summary(mod)
#>
#> Call:
#> lm(formula = body_mass_g ~ bill_length_mm + species, data = drop_na(penguins))
#>
#> Residuals:
#> Min 1Q Median 3Q Max
#> -860.77 -244.79 4.36 215.73 1075.25
#>
#> Coefficients:
#> Estimate Std. Error t value Pr(>|t|)
#> (Intercept) 200.453 271.646 0.738 0.461
#> bill_length_mm 90.298 6.951 12.991 < 2e-16 ***
#> speciesChinstrap -876.942 88.744 -9.882 < 2e-16 ***
#> speciesGentoo 596.702 76.429 7.807 7.88e-14 ***
#> ---
#> Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
#>
#> Residual standard error: 375.2 on 329 degrees of freedom
#> Multiple R-squared: 0.7848, Adjusted R-squared: 0.7829
#> F-statistic: 400 on 3 and 329 DF, p-value: < 2.2e-16Reading the output:
Estimate— change in y per 1-unit increase in xPr(>|t|)— p-value: is the coefficient ≠ 0?Adjusted R²— proportion of variance explainedF-statistic— overall model significance
→ broom wraps this into tidy tibbles (next slide)
Recap: broom — tidying model outputs ![]()
broom converts messy model objects into tidy tibbles. Three core functions:
| Function | Returns | Use it when you want to … |
|---|---|---|
tidy() |
One row per model term | Inspect coefficients, p-values, confidence intervals |
glance() |
One row per model | Compare overall fit (R², AIC, log-likelihood, …) |
augment() |
One row per observation | Add predictions, residuals, influence stats to data |
simple_mod <- lm(body_mass_g ~ bill_length_mm + species, data = drop_na(penguins))
tidy(simple_mod) # coefficients table
#> # A tibble: 4 × 5
#> term estimate std.error statistic p.value
#> <chr> <dbl> <dbl> <dbl> <dbl>
#> 1 (Intercept) 200. 272. 0.738 4.61e- 1
#> 2 bill_length_mm 90.3 6.95 13.0 1.97e-31
#> 3 speciesChinstrap -877. 88.7 -9.88 2.46e-20
#> 4 speciesGentoo 597. 76.4 7.81 7.88e-14Recap: broom in practice
glance(simple_mod) # model-level fit statistics
#> # A tibble: 1 × 12
#> r.squared adj.r.squared sigma statistic p.value df logLik AIC BIC
#> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
#> 1 0.785 0.783 375. 400. 2.22e-109 3 -2444. 4899. 4918.
#> # ℹ 3 more variables: deviance <dbl>, df.residual <int>, nobs <int>
augment(simple_mod) |>
select(body_mass_g, .fitted, .resid) |>
head(5)
#> # A tibble: 5 × 3
#> body_mass_g .fitted .resid
#> <int> <dbl> <dbl>
#> 1 3750 3731. 18.9
#> 2 3800 3767. 32.8
#> 3 3250 3839. -589.
#> 4 3450 3514. -64.4
#> 5 3650 3749. -99.1Note
augment() on a workflow (next slide) works the same way — it just accepts new_data = so you can apply the fitted model to held-out data. The column names (.fitted, .pred_class, .pred_*, .resid) are always predictable.
Predict with your model ![]()
How do you use your new model?
augment()will return the dataset with predictions and residuals added.
#> # A tibble: 333 × 9
#> body_mass_g bill_length_mm species .fitted .resid .hat .sigma .cooksd
#> <int> <dbl> <fct> <dbl> <dbl> <dbl> <dbl> <dbl>
#> 1 3750 39.1 Adelie 3731. 18.9 0.00688 376. 0.00000443
#> 2 3800 39.5 Adelie 3767. 32.8 0.00701 376. 0.0000136
#> 3 3250 40.3 Adelie 3839. -589. 0.00760 374. 0.00476
#> 4 3450 36.7 Adelie 3514. -64.4 0.00840 376. 0.0000629
#> 5 3650 39.3 Adelie 3749. -99.1 0.00693 376. 0.000123
#> 6 3625 38.9 Adelie 3713. -88.0 0.00685 376. 0.0000956
#> 7 4675 39.2 Adelie 3740. 935. 0.00690 372. 0.0109
#> 8 3200 41.1 Adelie 3912. -712. 0.00863 374. 0.00790
#> 9 3800 38.6 Adelie 3686. 114. 0.00687 376. 0.000161
#> 10 4400 34.6 Adelie 3325. 1075. 0.0130 371. 0.0273
#> # ℹ 323 more rows
#> # ℹ 1 more variable: .std.resid <dbl>Our typical process will look like this:
- Read in raw data (
readr,dplyr::*_join) (single or multi table)
- Prepare the data (EDA/mutate/summarize/clean, etc)
- This is “Feature Engineering”
- Once we’ve established our features (rows), we decide how to “spend” them …
For machine learning, we typically split data into training and test sets:
The training set is used to estimate model parameters.
Spending too much data in training prevents us from computing a good assessment of model performance.
The test set is used as an independent assessment of performance.
Spending too much data in testing prevents us from computing good model parameter estimates.
Exploratory Data Analysis
What is EDA?
- Exploratory Data Analysis (EDA) is an approach to analyzing datasets to summarize their key characteristics using visualizations and statistical techniques.
- It uncovers patterns, anomalies, and relationships — and guides data cleaning, feature selection, and model choice.
- Today we focus on EDA from a statistical perspective: understanding data properties needed to support modeling assumptions.
Why is EDA Important?
✅ Detects data quality issues – Missing values, outliers, inconsistencies.
✅ Identifies patterns & trends – Helps understand distributions and relationships.
✅ Reduces bias & assumptions – Ensures models are based on well-understood data.
✅ Supports feature engineering – Helps select or create the best features for modeling.
✅ Guides modeling choices – Informs whether linear, nonlinear, or other methods are suitable.
EDA Checklist (Peng 2016)
- Develop a question — E.g., “Is there a relationship between tree canopy and flood risk?”
- Read in data and check the structure — dimensions, variable types, factors vs. characters
- Summarize the data — missing values, ranges, means/medians, category counts
- Look at top and bottom —
head(),tail(),skimr::skim()for anomalies - Answer your question with descriptive measures — distributions, central tendency, variation
- Follow up with additional questions — EDA is iterative
Key Steps in EDA
1️⃣ Understand Data Structure – Check types, dimensions, missing values. glimpse(), str(), summary(), vis_dat()
2️⃣ Summary Statistics – Mean, median, variance, skewness, kurtosis. skimr::skim(), summary(), table()
3️⃣ Visualizations – Histograms, box plots, scatter plots, correlation matrices. ggplot2, ggpubr, visdat
4️⃣ Identify Data Issues – Outliers, missing data, inconsistencies. vis_dat(), IQR / Z-score methods
5️⃣ Transformations & Feature Engineering – Scaling, encoding, derived features. recipes, tidymodels
Common Pitfalls in EDA
⚠ Ignoring domain context – Statistical insights must align with real-world meaning.
⚠ Over-reliance on visualizations – Needs statistical validation.
⚠ Dismissing small patterns – Some signals may be weak but significant.
⚠ Not documenting findings – EDA should inform later modeling stages.
Understanding Data Structure
glimpse(penguins)
#> Rows: 344
#> Columns: 8
#> $ species <fct> Adelie, Adelie, Adelie, Adelie, Adelie, Adelie, Adel…
#> $ island <fct> Torgersen, Torgersen, Torgersen, Torgersen, Torgerse…
#> $ bill_length_mm <dbl> 39.1, 39.5, 40.3, NA, 36.7, 39.3, 38.9, 39.2, 34.1, …
#> $ bill_depth_mm <dbl> 18.7, 17.4, 18.0, NA, 19.3, 20.6, 17.8, 19.6, 18.1, …
#> $ flipper_length_mm <int> 181, 186, 195, NA, 193, 190, 181, 195, 193, 190, 186…
#> $ body_mass_g <int> 3750, 3800, 3250, NA, 3450, 3650, 3625, 4675, 3475, …
#> $ sex <fct> male, female, female, NA, female, male, female, male…
#> $ year <int> 2007, 2007, 2007, 2007, 2007, 2007, 2007, 2007, 2007…
nrow(penguins)
#> [1] 344Summary Stats with skimr
| Name | penguins |
| Number of rows | 344 |
| Number of columns | 8 |
| _______________________ | |
| Column type frequency: | |
| numeric | 1 |
| ________________________ | |
| Group variables | None |
Variable type: numeric
| skim_variable | n_missing | complete_rate | mean | sd | p0 | p25 | p50 | p75 | p100 | hist |
|---|---|---|---|---|---|---|---|---|---|---|
| body_mass_g | 2 | 0.99 | 4201.75 | 801.95 | 2700 | 3550 | 4050 | 4750 | 6300 | ▃▇▆▃▂ |
Central Tendency & Variability
Central Tendency:
- Mean: Average of all values. Best for symmetric distributions without outliers.
- Median: Middle value when sorted. Robust to outliers.
- Mode: Most frequent value. Useful for categorical data.
Variability:
- Range: Max − Min.
- Variance / SD: Spread around the mean.
- IQR: Range of the middle 50%; robust to outliers.
Overview of ggpubr
ggpubrwrapsggplot2to produce publication-ready statistical plots with minimal code.✅ Simplified syntax — no deep ggplot2 knowledge required
✅ Statistical plot types — box plots, violin, scatter, density, bar, dot
✅ Built-in tests —
stat_compare_means()adds p-values directly to plots✅ Grouped data — easy faceting, p-value annotation, confidence intervals
✅ Multi-plot layout —
ggarrange()combines plots in a grid
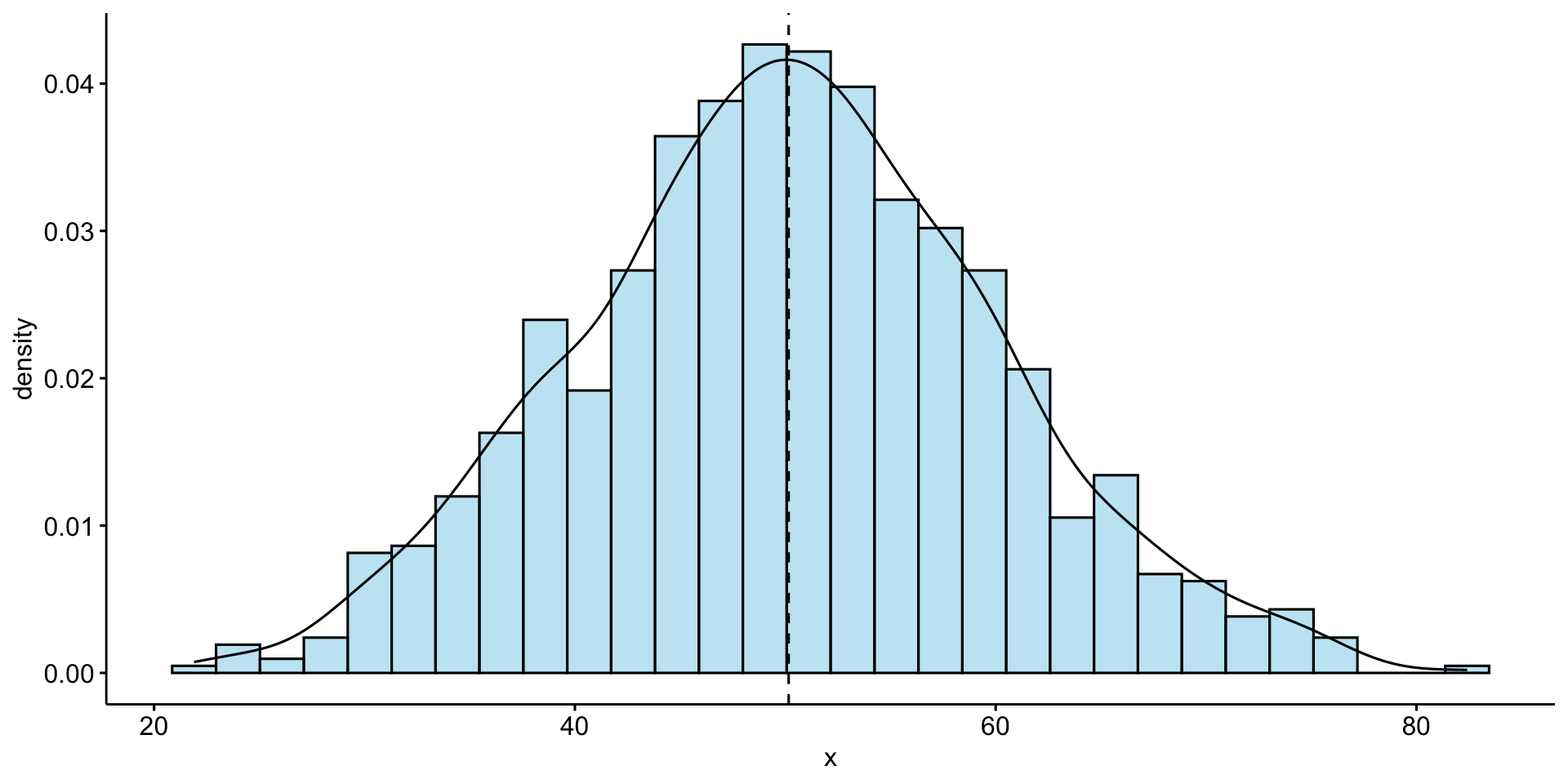
Univariate Analysis – Distribution of Bill Length
Univariate analysis focuses on a single variable at a time.
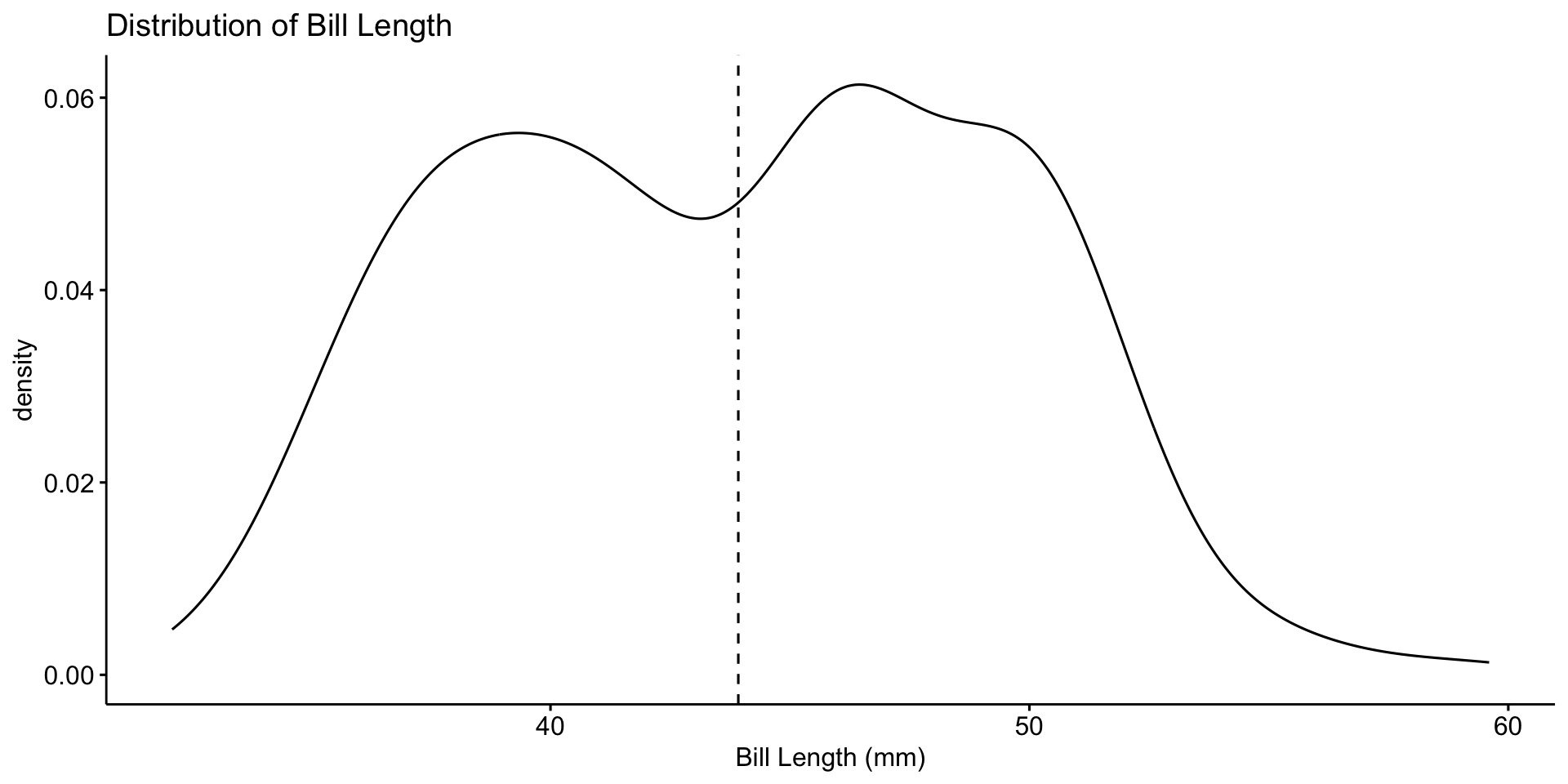
📌 Most bill lengths are between 35–55 mm. Some outliers exist.
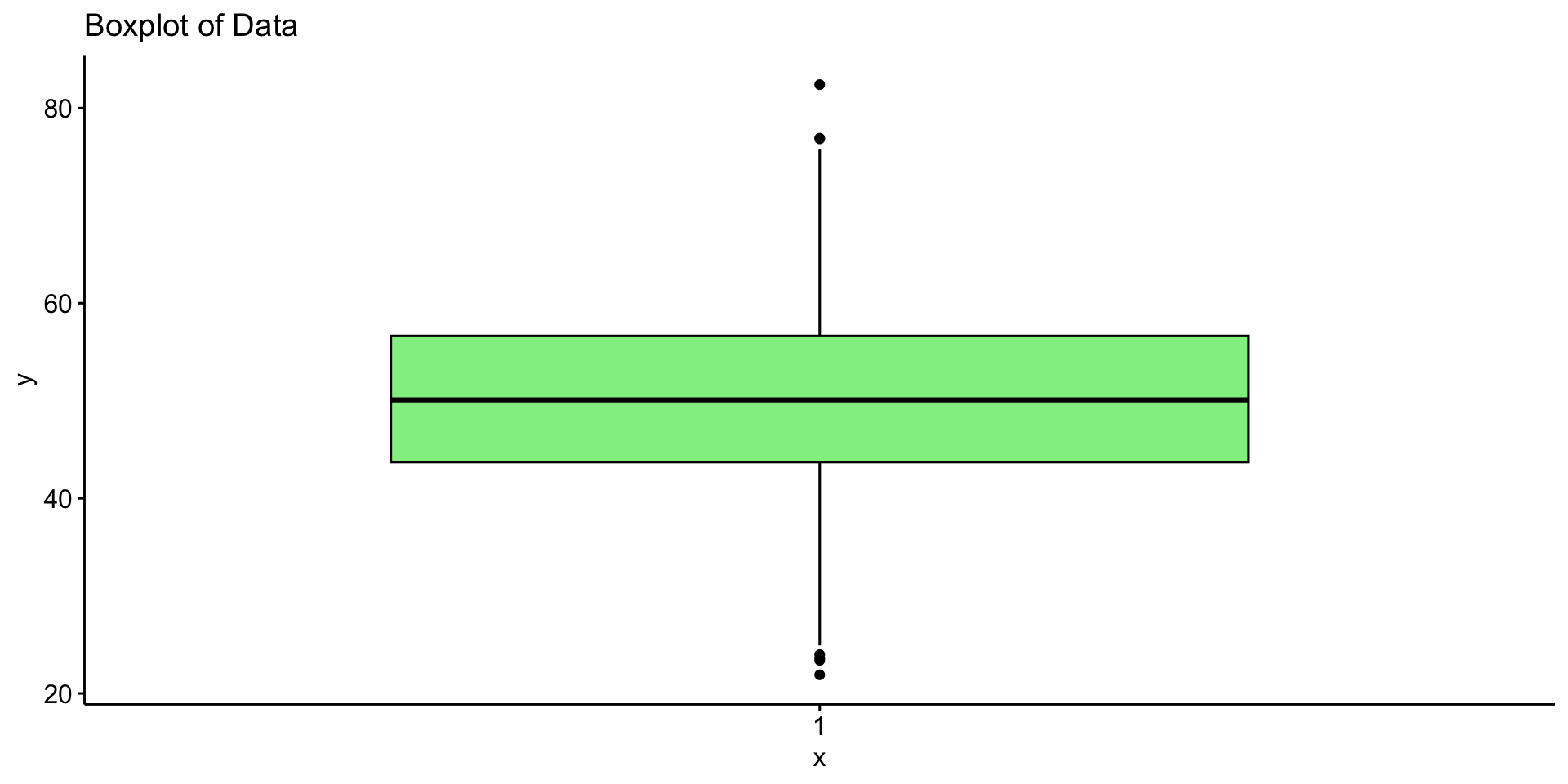
Boxplot for Outlier Detection
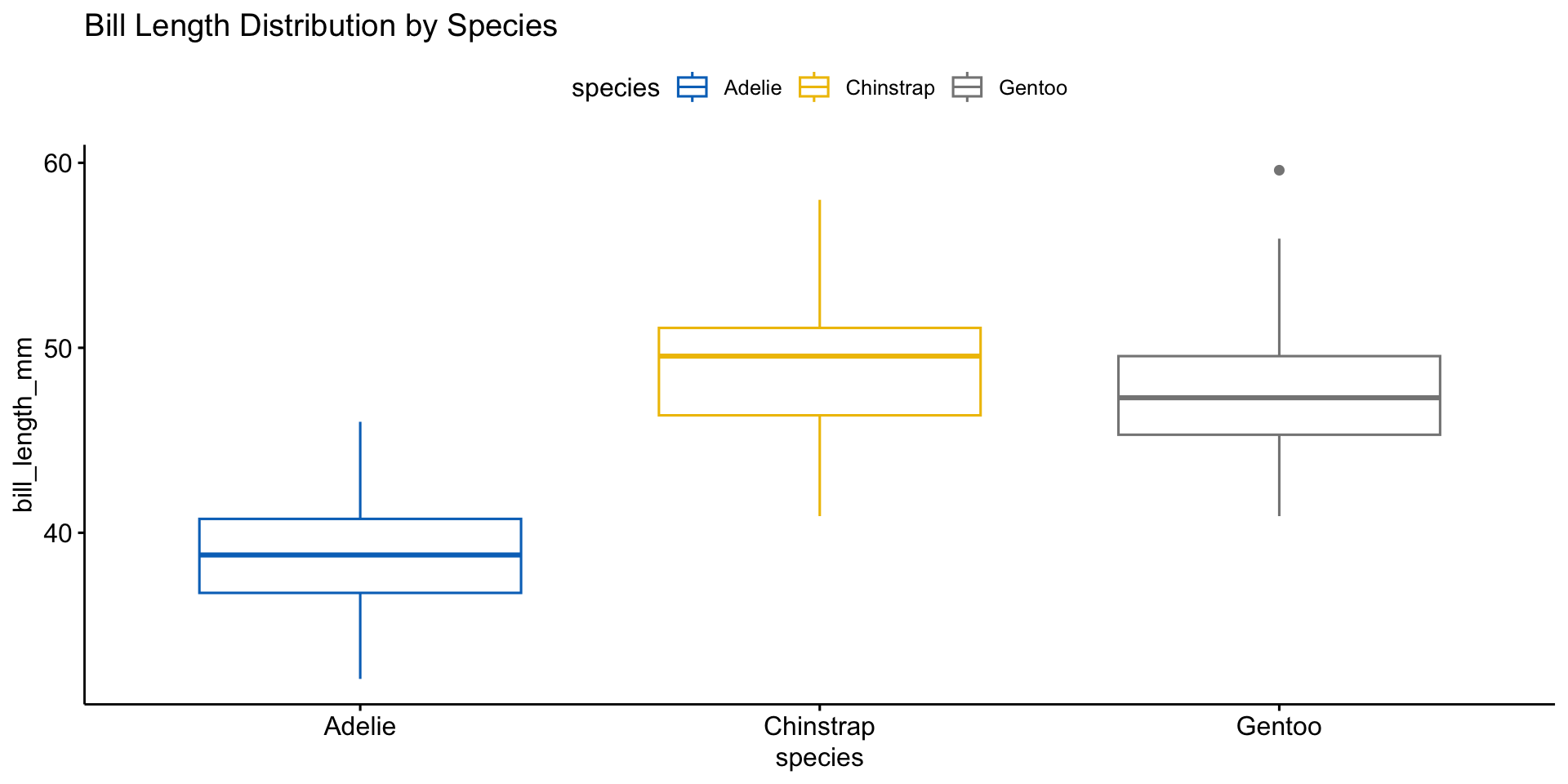
📌 Different species have distinct distributions. Outliers may need further investigation.
Bivariate Analysis
| Variable Types | Plot Type |
|---|---|
| Categorical + Categorical | Bar plot (stacked or side-by-side) |
| Categorical + Continuous | Box plot; bar plot with summary stats |
| Continuous + Continuous | Scatter plot with regression line |
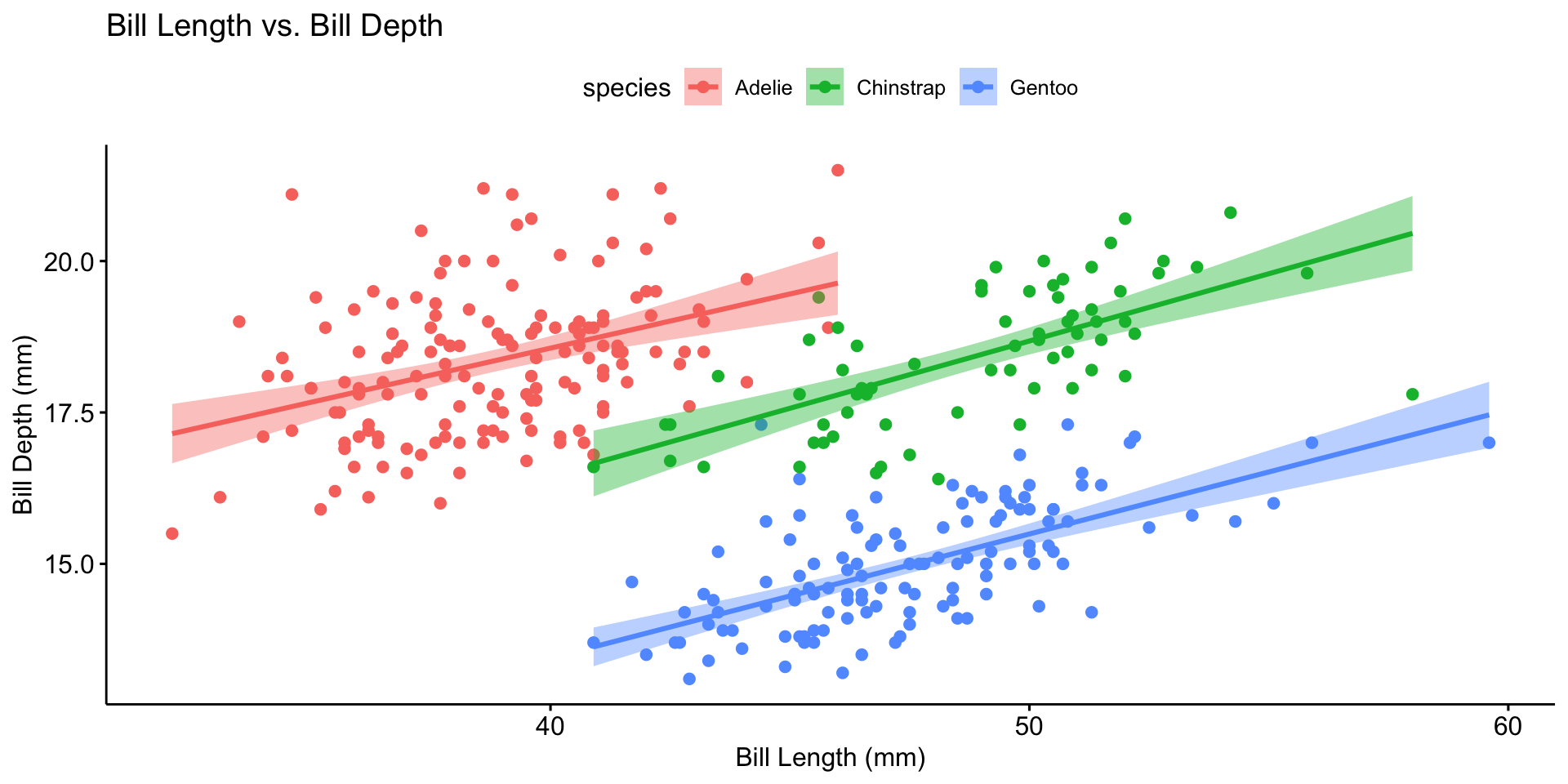
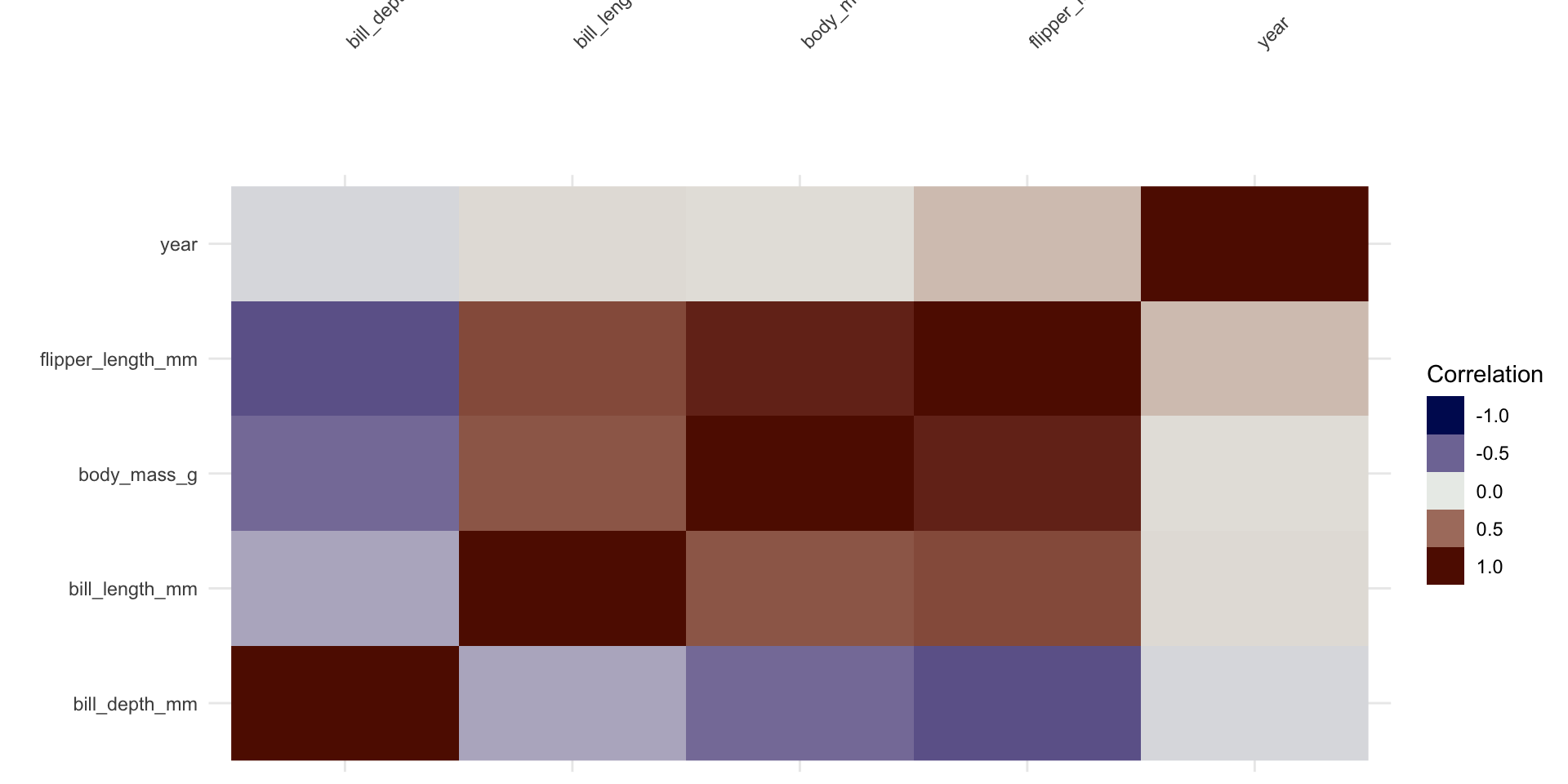
Multivariate Analysis
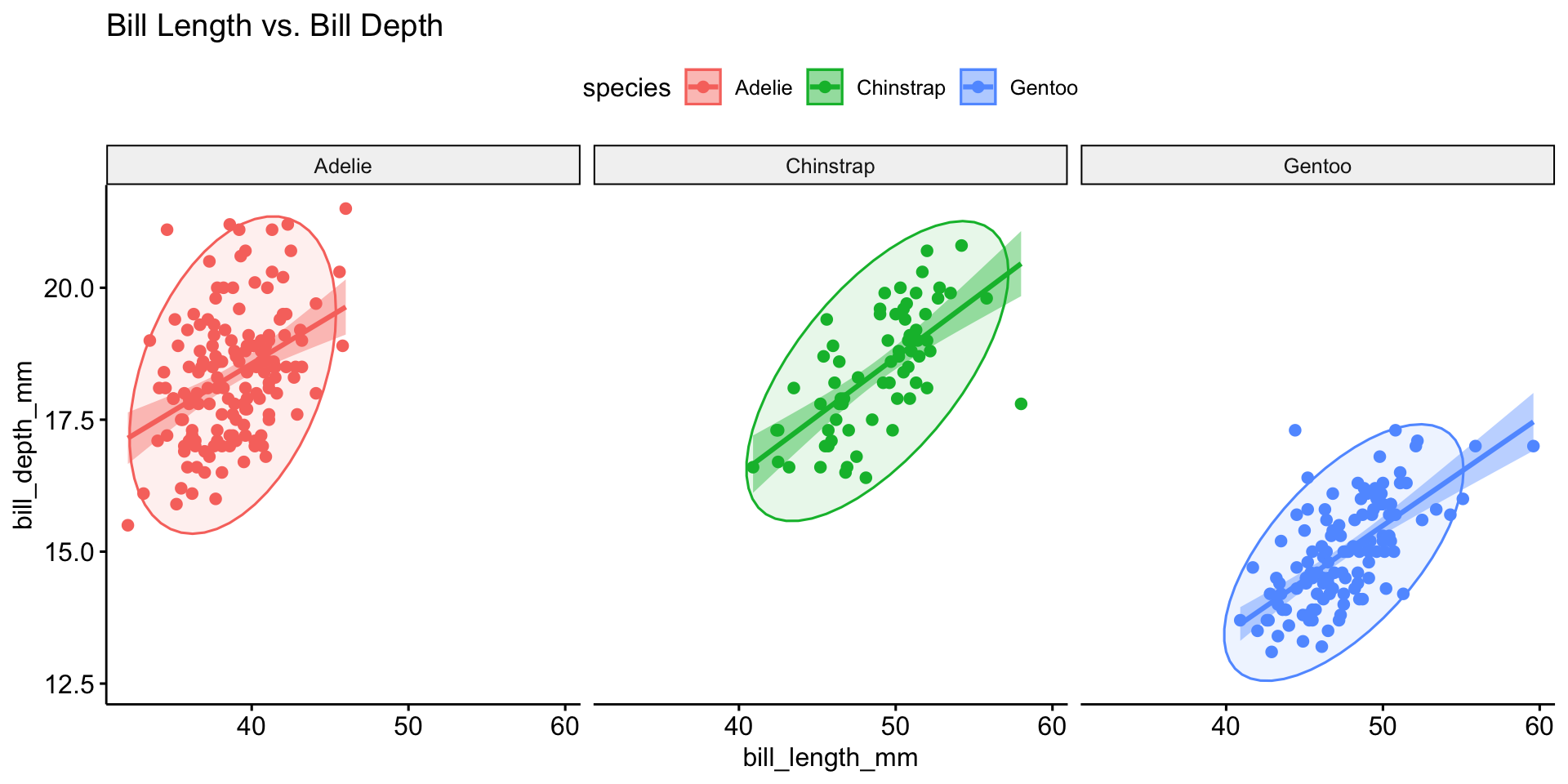
Correlation Analysis
Identify Data Issues
Duplicates:
Missing Data:
Outlier Detection: IQR, Z-scores, or clustering-based methods.
Data Normality
Many statistical tests and models rely on the assumption that data is normally distributed.
What is the Central Limit Theorem?
- The CLT states that the sampling distribution of the sample mean approaches a normal distribution as sample size increases, regardless of the original distribution.
- This justifies using normal-based inference even when raw data isn’t normal.
CLT in Action: Uniform Distribution
At n = 2, this may look normal, but the true distribution is triangular—CLT describes convergence, not instant normality
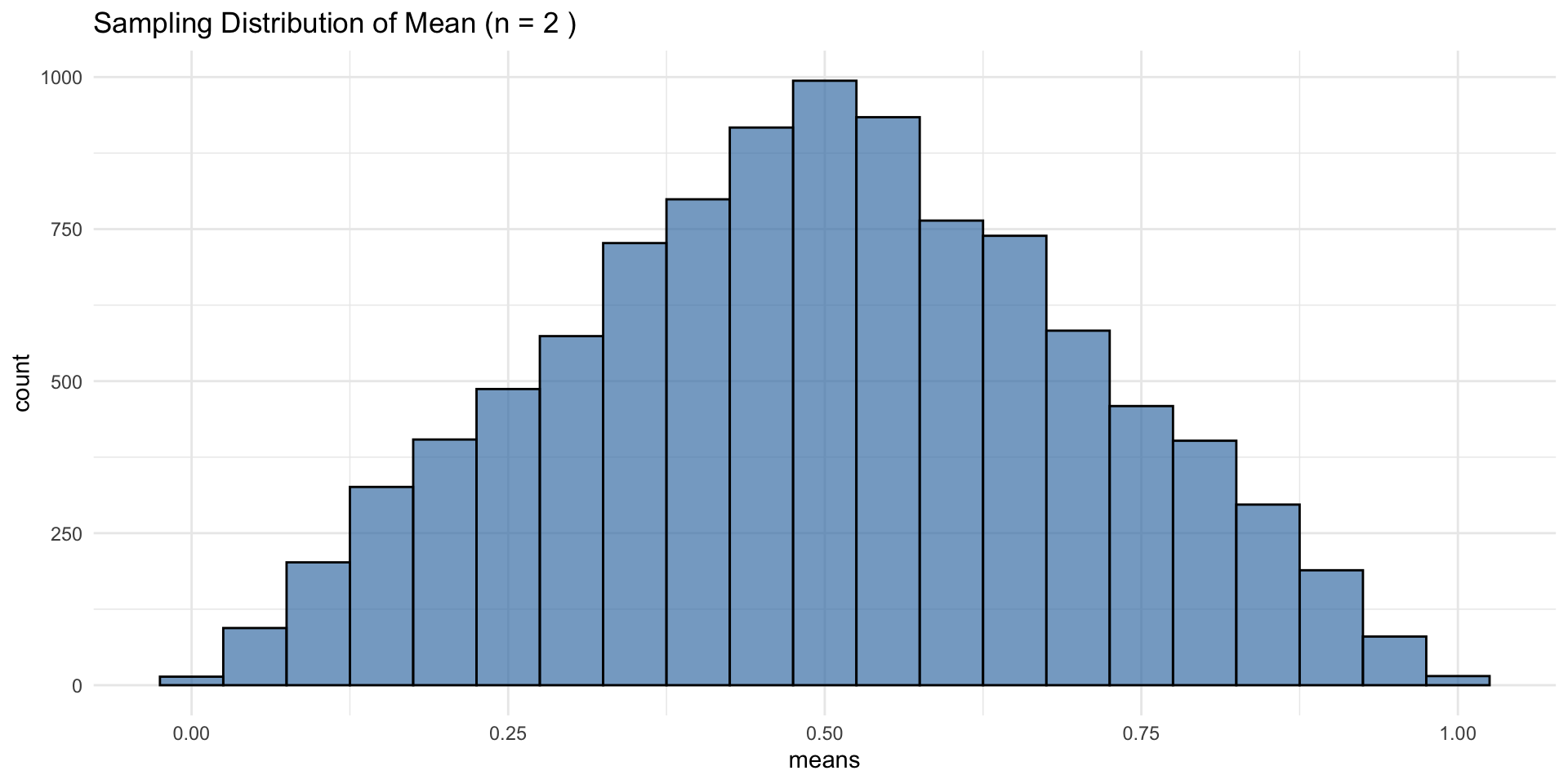
CLT: Increasing Sample Size
As n increases, the distribution of sample means becomes normal even though the original data was uniform.
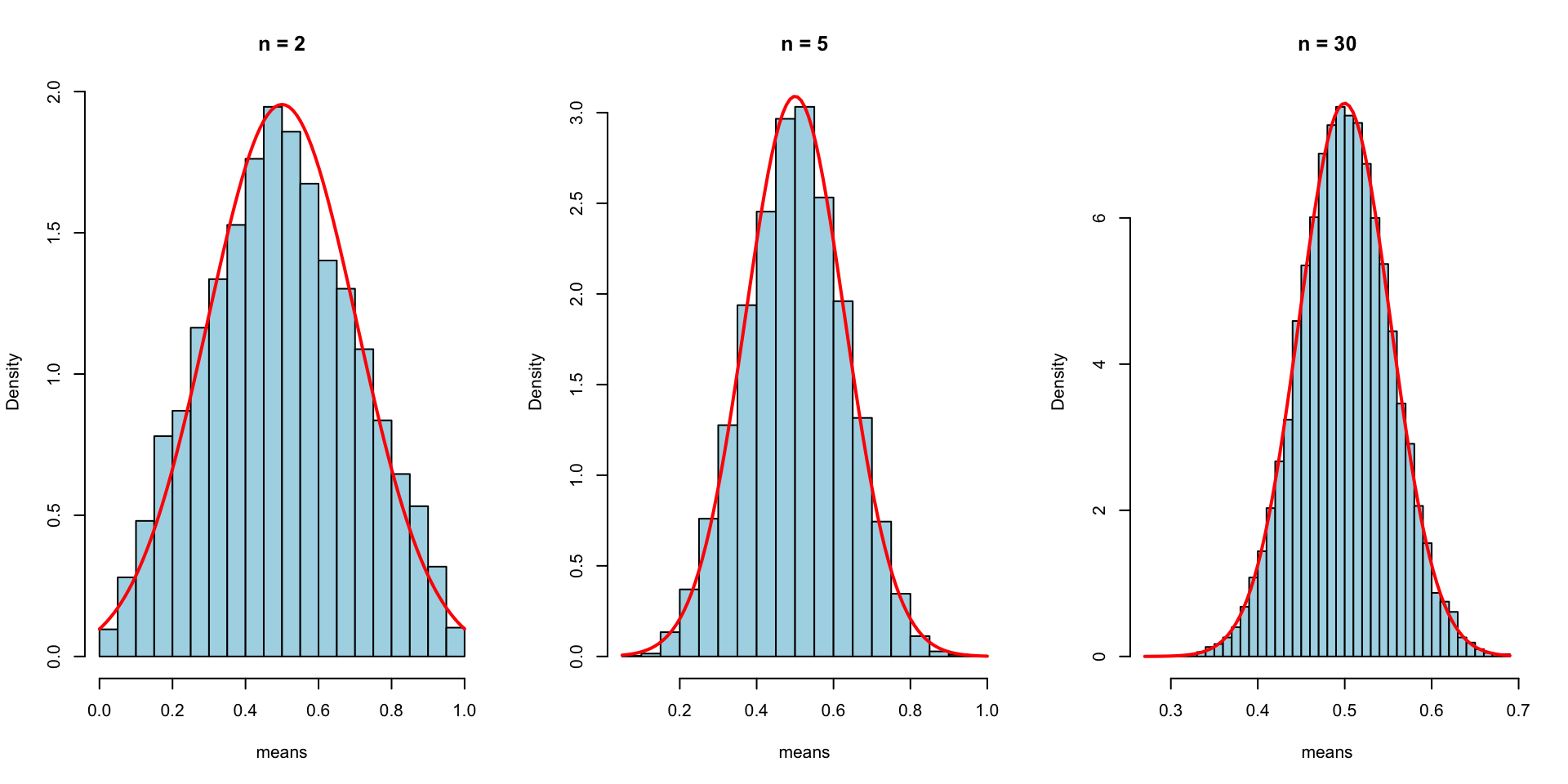
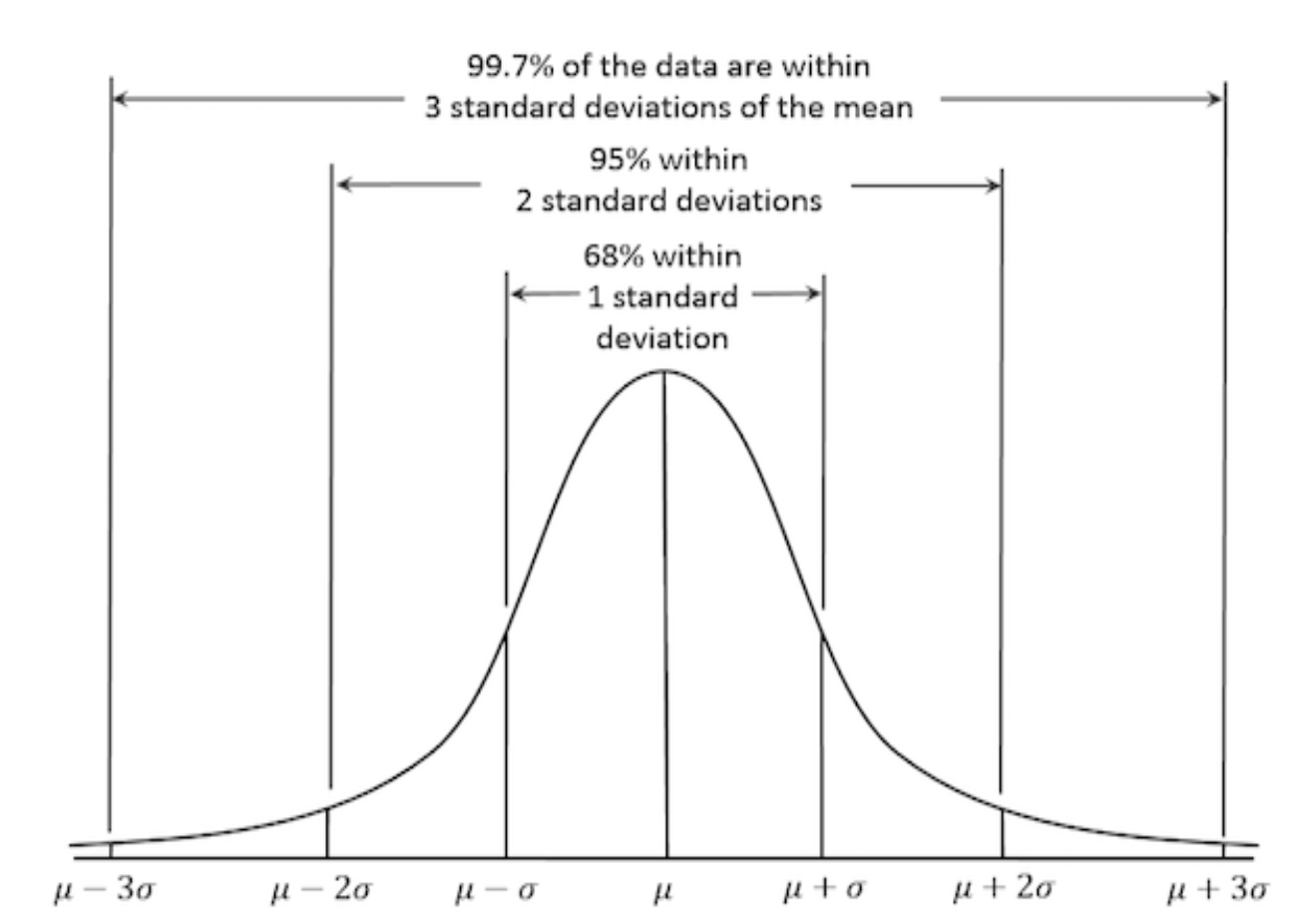
Properties of a Normal Distribution
Why Normality Matters
Many statistical techniques assume normality:
- t-tests / ANOVA — compare means between groups
- Linear regression (
lm) — residuals should be normally distributed - PCA — performs best with normally distributed features
- Confidence intervals — normality ensures accurate estimation
When data deviates from normality, these methods may produce inaccurate results.
Assessing Normality: Visual Methods
Checking Normality with Q-Q Plots
- Points along the 45-degree line → normally distributed.
- Deviations suggest skewness or outliers.
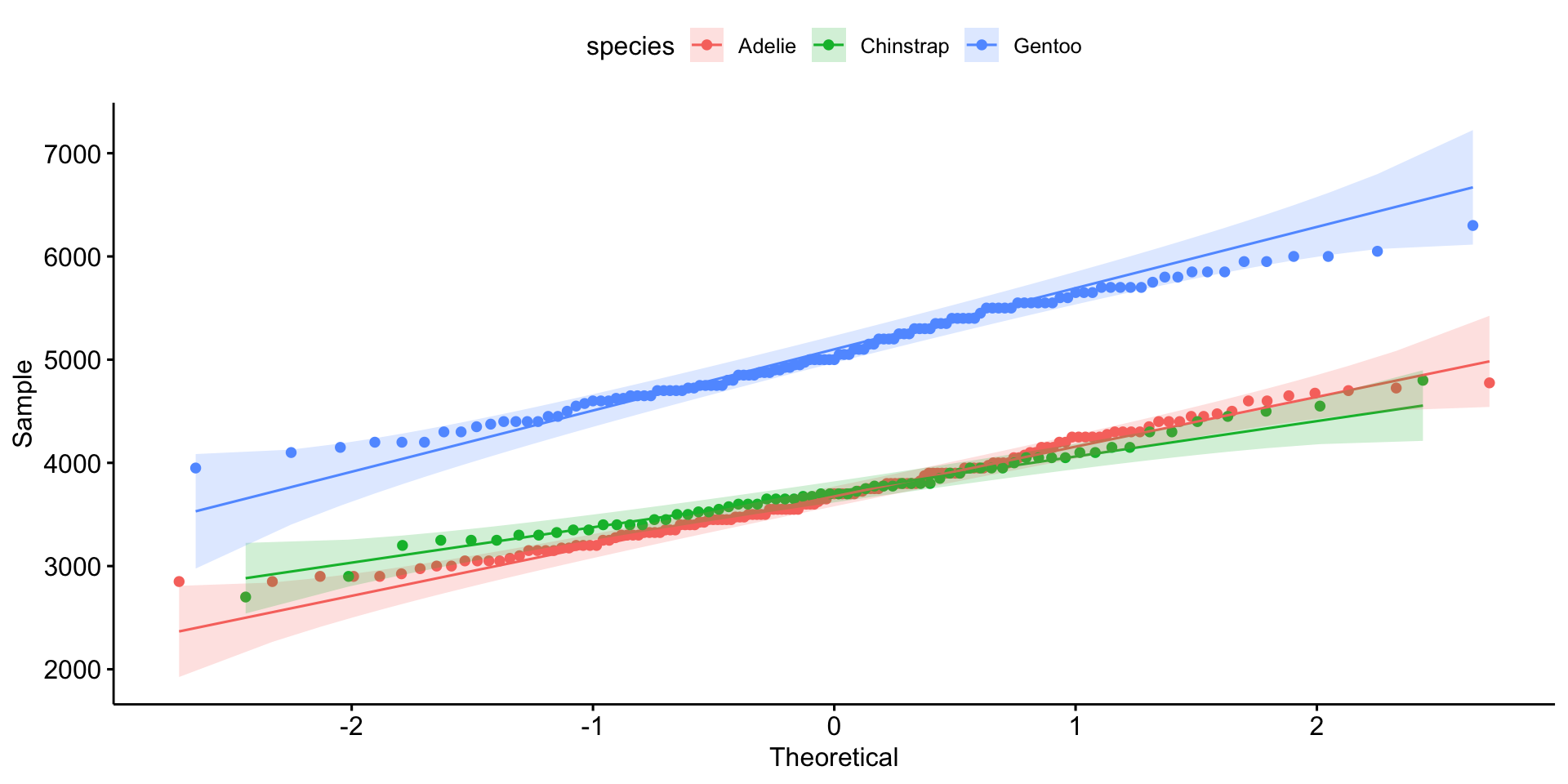
Statistical Tests for Normality
| Test | Strengths | Weaknesses |
|---|---|---|
| Shapiro-Wilk | Sensitive; good for small samples | Too sensitive for large n |
| Kolmogorov-Smirnov | Works for any sample size | Less powerful for small n |
| Anderson-Darling | Emphasizes tails | Sensitive to sample size |
| Jarque-Bera | Tests skewness + kurtosis | Less reliable for small n |
H₀ in all tests: data is normally distributed. Low p-value → reject H₀.
Normality Tests in R
nest/map Pattern for Per-Group Testing
penguins |>
drop_na() |>
nest(data = -species) |>
mutate(test = map(data, ~shapiro.test(.x$body_mass_g)),
glance_shapiro = map(test, broom::glance)) |>
unnest(glance_shapiro)
#> # A tibble: 3 × 6
#> species data test statistic p.value method
#> <fct> <list> <list> <dbl> <dbl> <chr>
#> 1 Adelie <tibble [146 × 7]> <htest> 0.981 0.0423 Shapiro-Wilk normality…
#> 2 Gentoo <tibble [119 × 7]> <htest> 0.986 0.261 Shapiro-Wilk normality…
#> 3 Chinstrap <tibble [68 × 7]> <htest> 0.984 0.561 Shapiro-Wilk normality…Dealing with Non-Normal Data
Option 1 — Transformations:
| Transformation | When to Use |
|---|---|
| Log | Right-skewed, positive values |
| Square Root | Moderate right-skew, count data |
| Box-Cox | Unknown transformation needed |
| Yeo-Johnson | Like Box-Cox but works on negative values |
Option 2 — Nonparametric Methods
Use when data is not normally distributed:
# Spearman correlation (instead of Pearson)
cor.test(airquality$Ozone, airquality$Temp, method = "spearman")
#>
#> Spearman's rank correlation rho
#>
#> data: airquality$Ozone and airquality$Temp
#> S = 58778, p-value < 2.2e-16
#> alternative hypothesis: true rho is not equal to 0
#> sample estimates:
#> rho
#> 0.774043Option 3 — Bootstrapping
Provides robust estimates without assuming normality:
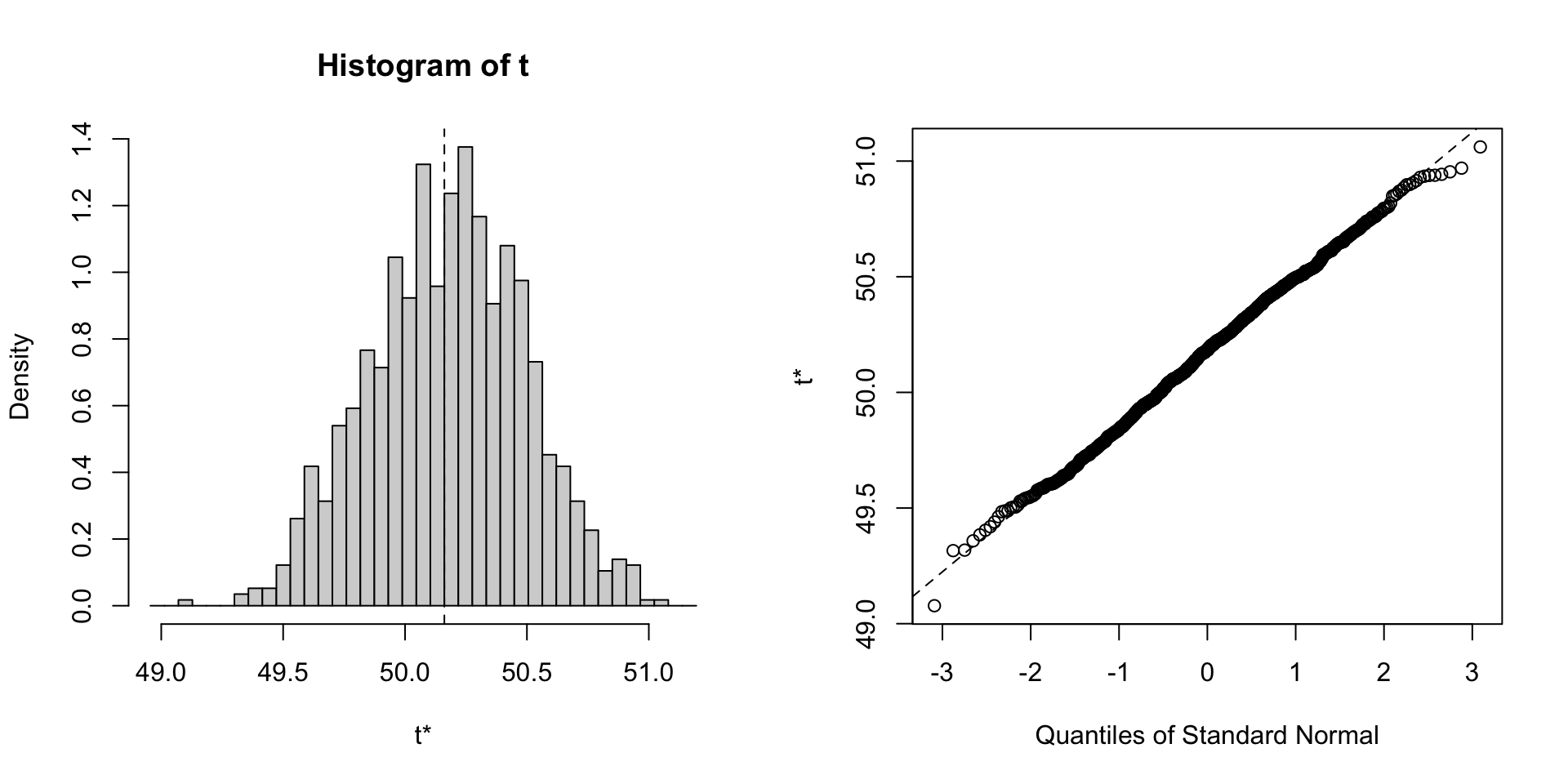
Skewness
Skewness measures the asymmetry of a distribution:
- Positive skew: Longer right tail
- Negative skew: Longer left tail
- Symmetric: No skew (normal)
D’Agostino test for skewness:
On to modeling!
Now that the data is cleaned, understood, and modified, we can move on to modeling.
Data Budget
In any modeling effort, it’s crucial to evaluate the performance of a model using different validation techniques to ensure a model can generalize to unseen data.
But data is limited even in the age of “big data”.
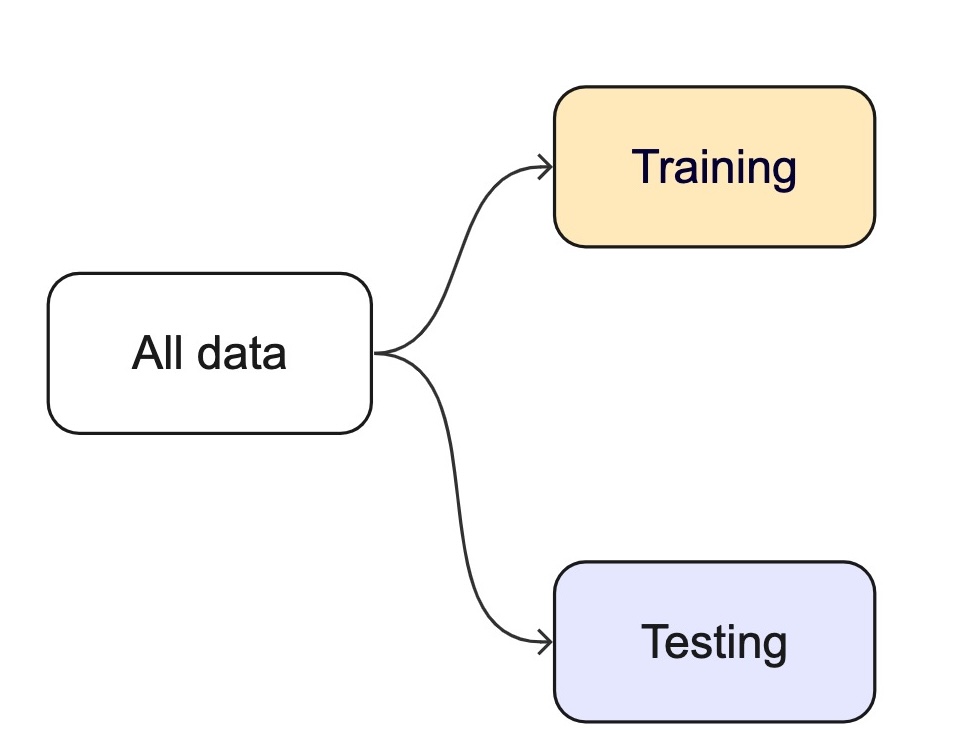
How we split data
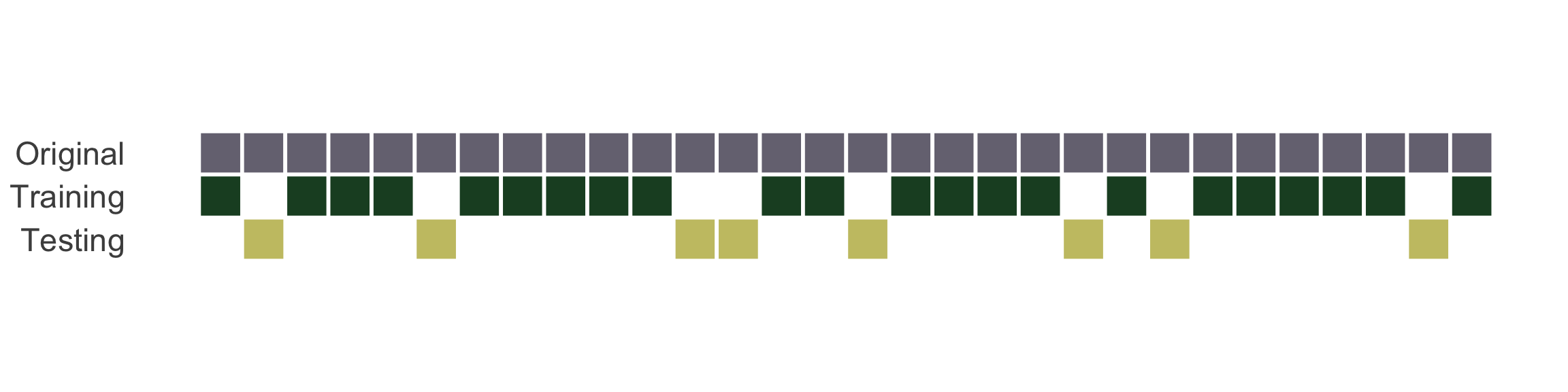

Splitting can be handled in many of ways. Typically, we base it off of a “hold out” percentage (e.g. 20%)
These hold out cases are extracted randomly from the data set. (remember seeds?)
- The training set is usually the majority of the data and provides a sandbox for testing different modeling approaches.
The test set is held in reserve until one or two models are chosen.
The test set is then used as the final arbiter to determine the efficacy of the model.
Stratification
In many cases, there is a structure to the data that would inhibit impartial splitting (e.g. a class, a region, a species, a sex, etc)
Imagine you have a jar of M&Ms in different colors — red, blue, and green. You want to take some candies out to taste, but you want to make sure you get a fair mix of each color, not just grabbing a bunch of red ones by accident.
Stratified resampling is like making sure that if your jar is 50% red, 30% blue, and 20% green, then the handful of candies you take keeps the same balance.
In data science, we do the same thing when we pick samples from a dataset: we make sure that different groups (like categories of people, animals, or weather types) are still fairly represented!
Initial splits ![]()
- In
tidymodels, thersamplepackage provides functions for creating initial splits of data. - The
initial_split()function is used to create a single split of the data. - The
propargument defines the proportion of the data to be used for training. - The default is 0.75, which means 75% of the data will be used for training and 25% for testing.
Accessing the data: ![]()
- Once the data is split, we can access the training and testing data using the
training()andtesting()functions to extract the partitioned data from the full set:
penguins_train <- training(resample_split)
glimpse(penguins_train)
#> Rows: 275
#> Columns: 8
#> $ species <fct> Gentoo, Adelie, Gentoo, Chinstrap, Gentoo, Chinstrap…
#> $ island <fct> Biscoe, Torgersen, Biscoe, Dream, Biscoe, Dream, Dre…
#> $ bill_length_mm <dbl> 46.2, 43.1, 46.2, 45.9, 46.5, 48.1, 45.2, 55.9, 38.1…
#> $ bill_depth_mm <dbl> 14.9, 19.2, 14.4, 17.1, 13.5, 16.4, 17.8, 17.0, 17.0…
#> $ flipper_length_mm <int> 221, 197, 214, 190, 210, 199, 198, 228, 181, 195, 20…
#> $ body_mass_g <int> 5300, 3500, 4650, 3575, 4550, 3325, 3950, 5600, 3175…
#> $ sex <fct> male, male, NA, female, female, female, female, male…
#> $ year <int> 2008, 2009, 2008, 2007, 2007, 2009, 2007, 2009, 2009…penguins_test <- testing(resample_split)
glimpse(penguins_test)
#> Rows: 69
#> Columns: 8
#> $ species <fct> Adelie, Adelie, Adelie, Adelie, Adelie, Adelie, Adel…
#> $ island <fct> Torgersen, Torgersen, Torgersen, Biscoe, Biscoe, Bis…
#> $ bill_length_mm <dbl> NA, 39.3, 36.6, 35.9, 38.8, 37.9, 39.2, 39.6, 36.7, …
#> $ bill_depth_mm <dbl> NA, 20.6, 17.8, 19.2, 17.2, 18.6, 21.1, 17.2, 18.8, …
#> $ flipper_length_mm <int> NA, 190, 185, 189, 180, 172, 196, 196, 187, 205, 187…
#> $ body_mass_g <int> NA, 3650, 3700, 3800, 3800, 3150, 4150, 3550, 3800, …
#> $ sex <fct> NA, male, female, female, male, female, male, female…
#> $ year <int> 2007, 2007, 2007, 2007, 2007, 2007, 2007, 2008, 2008…Proportional Gaps — Before Stratification
Warning
Without stratification, random sampling can over- or under-represent groups. Compare the species proportions below — they don’t match.
# Dataset
(table(penguins$species) / nrow(penguins))
#>
#> Adelie Chinstrap Gentoo
#> 0.4418605 0.1976744 0.3604651
# Training Data
(table(penguins_train$species) / nrow(penguins_train))
#>
#> Adelie Chinstrap Gentoo
#> 0.4581818 0.1818182 0.3600000
# Testing Data
(table(penguins_test$species) / nrow(penguins_test))
#>
#> Adelie Chinstrap Gentoo
#> 0.3768116 0.2608696 0.3623188Stratification Example
A stratified random sample conducts a specified split within defined subsets of subsets, and then pools the results.
Only one column can be used to define a strata, but grouping/mutate operations can be used to create a new column that can be used as a strata (e.g. species/island)
In the case of the penguins data, we can stratify by species (think back to our nested/linear model approach)
This ensures that the training and testing sets have the same proportion of each species as the original data.
Proportional Alignment
# Check the proportions
# Dataset
table(penguins$species) / nrow(penguins)
#>
#> Adelie Chinstrap Gentoo
#> 0.4384384 0.2042042 0.3573574
# Training Data
table(train_strata$species) / nrow(train_strata)
#>
#> Adelie Chinstrap Gentoo
#> 0.4377358 0.2037736 0.3584906
# Testing Data
table(test_strata$species) / nrow(test_strata)
#>
#> Adelie Chinstrap Gentoo
#> 0.4411765 0.2058824 0.3529412Modeling
Once the data is split, we can decide what type of model to invoke.
Often, users simply pick a well known model type or class for a type of problem (Classic Conceptual Model)
We will learn more about model types and uses next week!
For now, just know that the training data is used to fit a model.
If we are certain about the model, we can use the test data to evaluate the model.

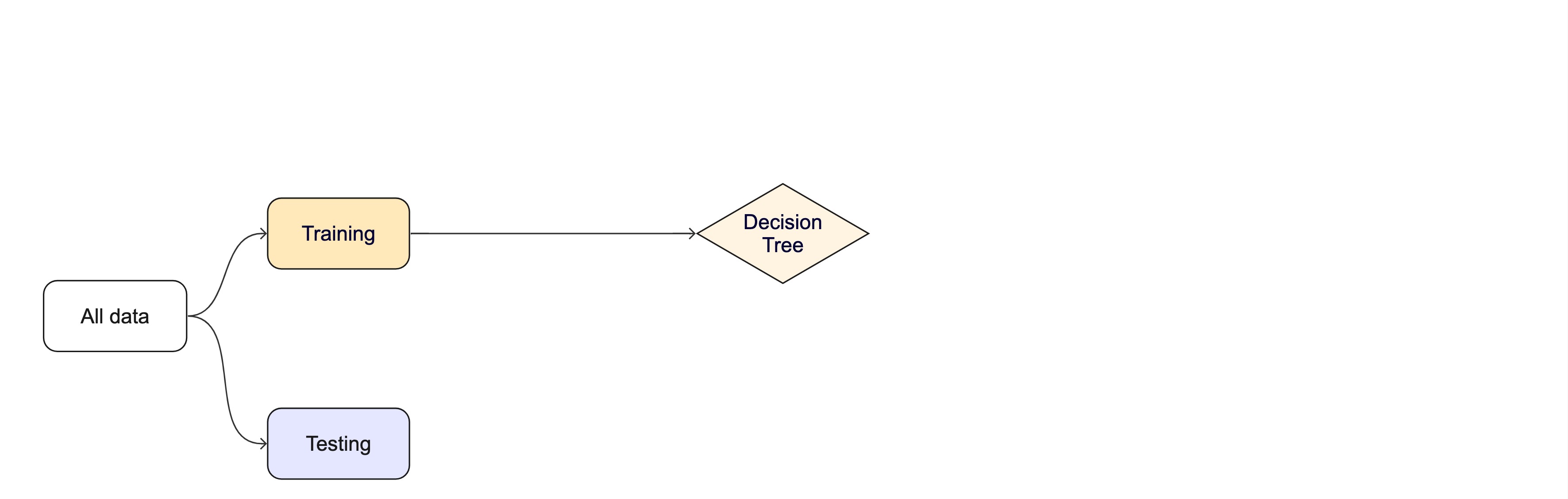
Modeling Options
But, there are many types of models, with different assumptions, behaviors, and qualities, that make some more applicable to a given dataset!
Most models have some type of parameterization, that can often be tuned.
Testing combinations of model and tuning parameters also requires some combination of
training/testingsplits.
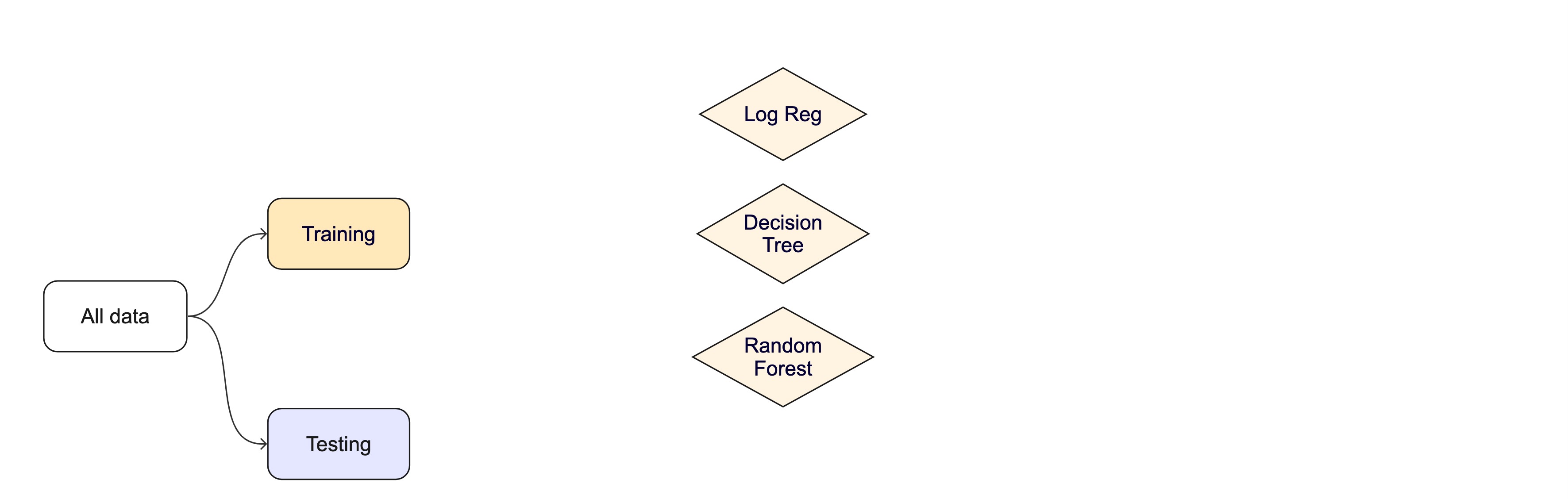
Modeling Options
What if we want to compare more models?
And/or more model configurations?
And we want to understand if these are important differences?
How can we use the training data to compare and evaluate different models? 🤔
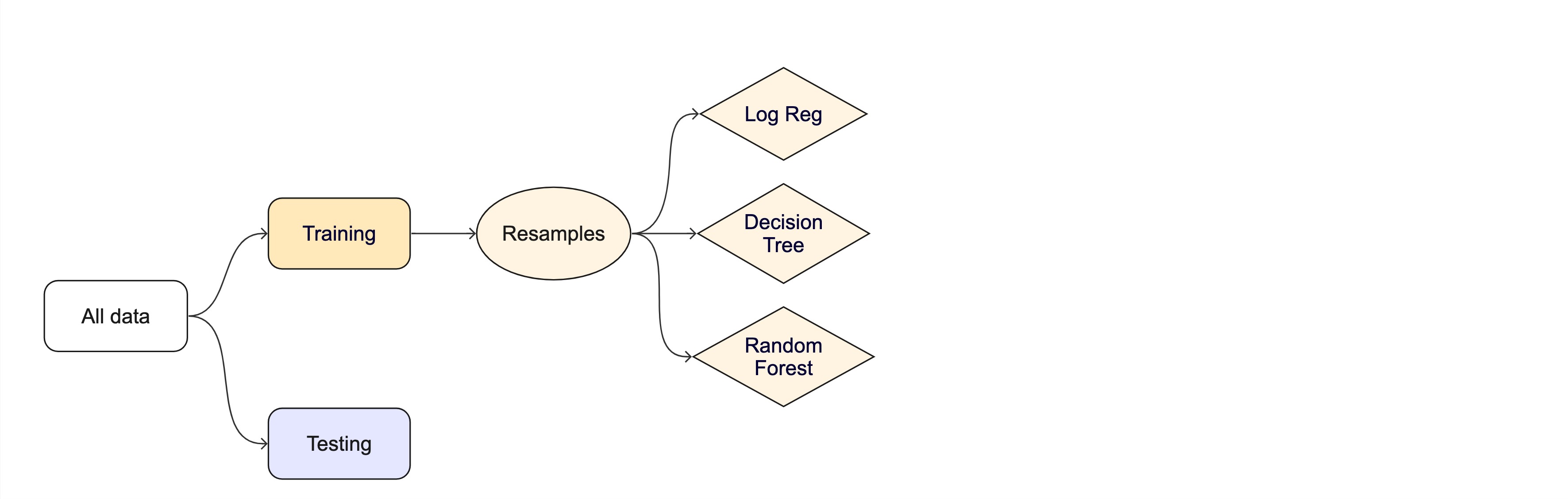
Resampling
Testing combinations of model and tuning parameters also requires some combination of
training/testingsplits.Resampling methods, such as cross-validation and bootstrapping, are empirical simulation systems that help facilitate this.
They create a series of data sets similar to the initial
training/testingsplit.In the first level of the diagram to the right, we first split the original data into
training/testingsets. Then, the training set is chosen for resampling.
Key Resampling Methods
Warning
Resampling is always used with the training set.
Resampling is a key step in model validation and assessment. The rsample package provides tools to create resampling strategies such as cross-validation, bootstrapping, and validation splits.
K-Fold Cross-Validation: The dataset is split into multiple “folds” (e.g., 5-fold or 10-fold), where each fold is used as a validation set while the remaining data is used for training.
Example (10-fold): On iteration 1, folds 2–10 become the training set and fold 1 is held out for assessment. On iteration 2, fold 2 is held out and folds 1, 3–10 are trained on. After 10 iterations every row has been predicted exactly once from a model it did not train on — and we average the 10 metric values to get a more stable estimate of skill.
- Bootstrap Resampling: Samples are drawn with replacement from the training dataset to generate multiple datasets, which are used to train and test the model.
- Monte Carlo Resampling (Repeated Train-Test Splits): Randomly splits data multiple times.
- Validation Split: Creates a single training and testing partition.
Cross-validation
Cross-validation
Cross-validation ![]()
penguins_train |> glimpse()
#> Rows: 275
#> Columns: 8
#> $ species <fct> Gentoo, Adelie, Gentoo, Chinstrap, Gentoo, Chinstrap…
#> $ island <fct> Biscoe, Torgersen, Biscoe, Dream, Biscoe, Dream, Dre…
#> $ bill_length_mm <dbl> 46.2, 43.1, 46.2, 45.9, 46.5, 48.1, 45.2, 55.9, 38.1…
#> $ bill_depth_mm <dbl> 14.9, 19.2, 14.4, 17.1, 13.5, 16.4, 17.8, 17.0, 17.0…
#> $ flipper_length_mm <int> 221, 197, 214, 190, 210, 199, 198, 228, 181, 195, 20…
#> $ body_mass_g <int> 5300, 3500, 4650, 3575, 4550, 3325, 3950, 5600, 3175…
#> $ sex <fct> male, male, NA, female, female, female, female, male…
#> $ year <int> 2008, 2009, 2008, 2007, 2007, 2009, 2007, 2009, 2009…nrow(penguins_train) * 1/10
#> [1] 27.5
vfold_cv(penguins_train, v = 10) # v = 10 is default
#> # 10-fold cross-validation
#> # A tibble: 10 × 2
#> splits id
#> <list> <chr>
#> 1 <split [247/28]> Fold01
#> 2 <split [247/28]> Fold02
#> 3 <split [247/28]> Fold03
#> 4 <split [247/28]> Fold04
#> 5 <split [247/28]> Fold05
#> 6 <split [248/27]> Fold06
#> 7 <split [248/27]> Fold07
#> 8 <split [248/27]> Fold08
#> 9 <split [248/27]> Fold09
#> 10 <split [248/27]> Fold10Cross-validation ![]()
What is in this?
Note
Here is another example of a list column enabling the storage of non-atomic types in tibble
Important
Set the seed when creating resamples
Alternate resampling schemes
Bootstrapping
Bootstrapping ![]()
set.seed(3214)
bootstraps(penguins_train)
#> # Bootstrap sampling
#> # A tibble: 25 × 2
#> splits id
#> <list> <chr>
#> 1 <split [275/96]> Bootstrap01
#> 2 <split [275/105]> Bootstrap02
#> 3 <split [275/108]> Bootstrap03
#> 4 <split [275/111]> Bootstrap04
#> 5 <split [275/102]> Bootstrap05
#> 6 <split [275/98]> Bootstrap06
#> 7 <split [275/92]> Bootstrap07
#> 8 <split [275/97]> Bootstrap08
#> 9 <split [275/100]> Bootstrap09
#> 10 <split [275/99]> Bootstrap10
#> # ℹ 15 more rowsMonte Carlo Cross-Validation ![]()
set.seed(322)
mc_cv(penguins_train, times = 10)
#> # Monte Carlo cross-validation (0.75/0.25) with 10 resamples
#> # A tibble: 10 × 2
#> splits id
#> <list> <chr>
#> 1 <split [206/69]> Resample01
#> 2 <split [206/69]> Resample02
#> 3 <split [206/69]> Resample03
#> 4 <split [206/69]> Resample04
#> 5 <split [206/69]> Resample05
#> 6 <split [206/69]> Resample06
#> 7 <split [206/69]> Resample07
#> 8 <split [206/69]> Resample08
#> 9 <split [206/69]> Resample09
#> 10 <split [206/69]> Resample10Validation set ![]()
A validation set is just another type of resample
The whole game - status update
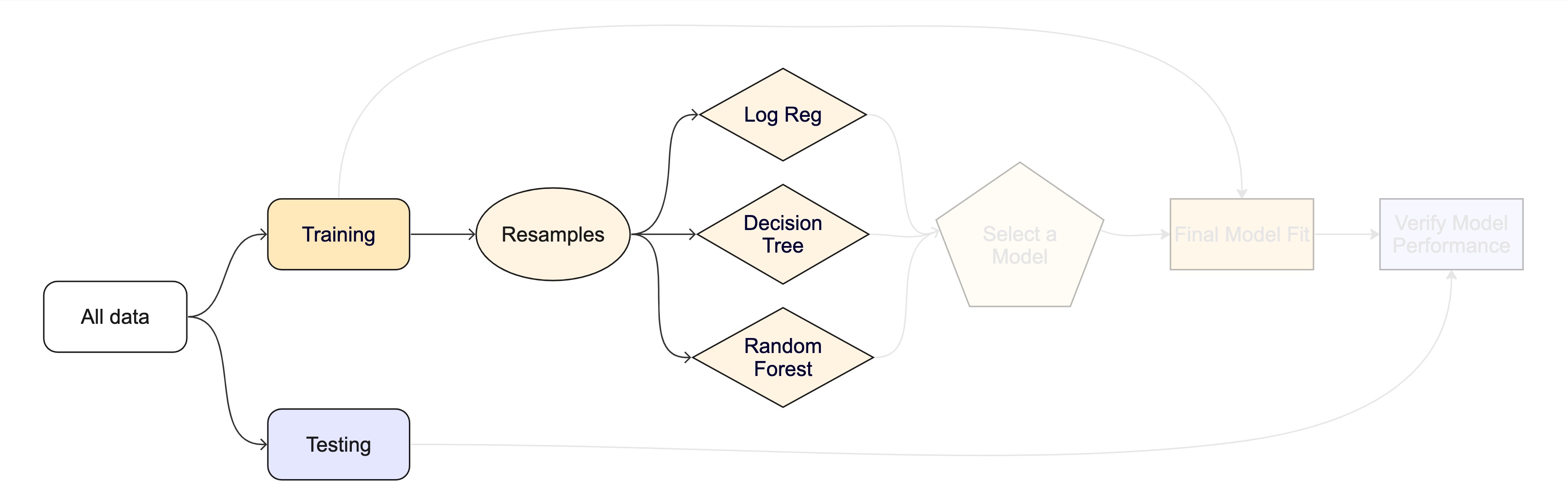
Unit Example
Can we predict if a plot of land is forested?
- Interested in classification models
- We have a dataset of 7,107 6000-acre hexagons in Washington state
- Each hexagon has a nominal outcome,
forested, with levels"Yes"and"No"
Data on forests in Washington
- The U.S. Forest Service maintains ML models to predict whether a plot of land is “forested.”
- This classification is important for all sorts of research, legislation, and land management purposes.
- Plots are typically remeasured every 10 years and this dataset contains the most recent measurement per plot.
- Type
?forestedto learn more about this dataset, including references.
Data on forests in Washington
One observation from each of 7,107 6000-acre hexagons in Washington state.
A nominal outcome,
forested, with levels"Yes"and"No", measured on-the-ground (expensive, slow).
18 remotely-sensed and easily-accessible predictors (cheap, fast, wall-to-wall coverage).
The ML framing: use cheap, abundant predictors to estimate an expensive ground-truth label so the Forest Service can predict forest cover without sending a crew to every hexagon.
Data splitting and spending
The initial split ![]()
Accessing the data ![]()
K-Fold Cross-Validation ![]()
# Load the dataset
set.seed(123)
# Create a 10-fold cross-validation object
(forested_folds <- vfold_cv(forested_train, v = 10))
#> # 10-fold cross-validation
#> # A tibble: 10 × 2
#> splits id
#> <list> <chr>
#> 1 <split [5116/569]> Fold01
#> 2 <split [5116/569]> Fold02
#> 3 <split [5116/569]> Fold03
#> 4 <split [5116/569]> Fold04
#> 5 <split [5116/569]> Fold05
#> 6 <split [5117/568]> Fold06
#> 7 <split [5117/568]> Fold07
#> 8 <split [5117/568]> Fold08
#> 9 <split [5117/568]> Fold09
#> 10 <split [5117/568]> Fold10How do you fit a model in R?
lmfor linear model <– The one we have looked at!glmnetfor regularized regressionkerasfor regression using TensorFlowstanfor Bayesian regression using Stansparkfor large data sets using sparkbruleefor regression using torch (PyTorch)
Challenge
- All of these models have different syntax and functions
- How do you keep track of all of them?
- How do you know which one to use?
- How would you compare them?
Comparing 3-5 of these models is a lot of work using functions for diverse packages.
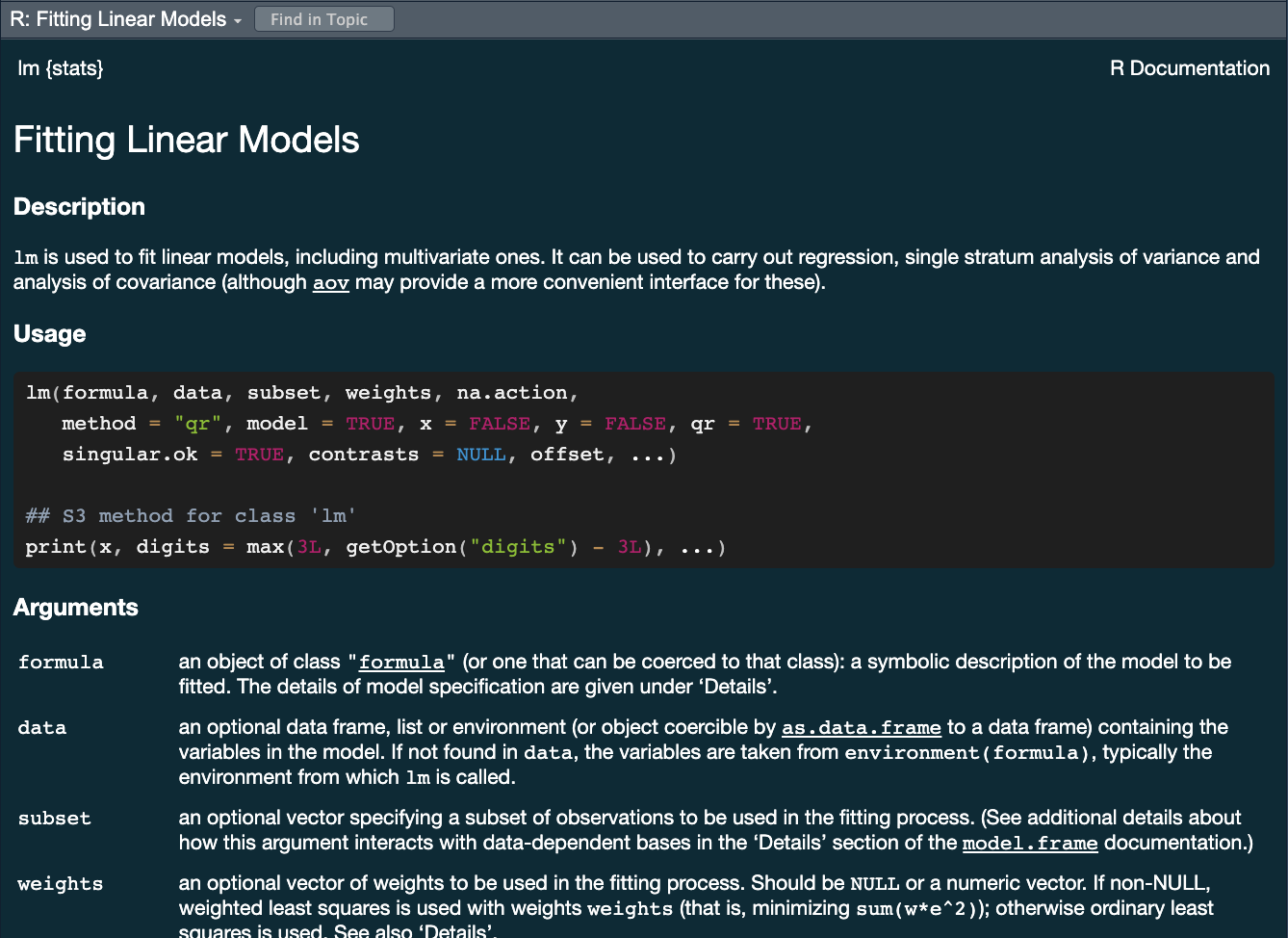
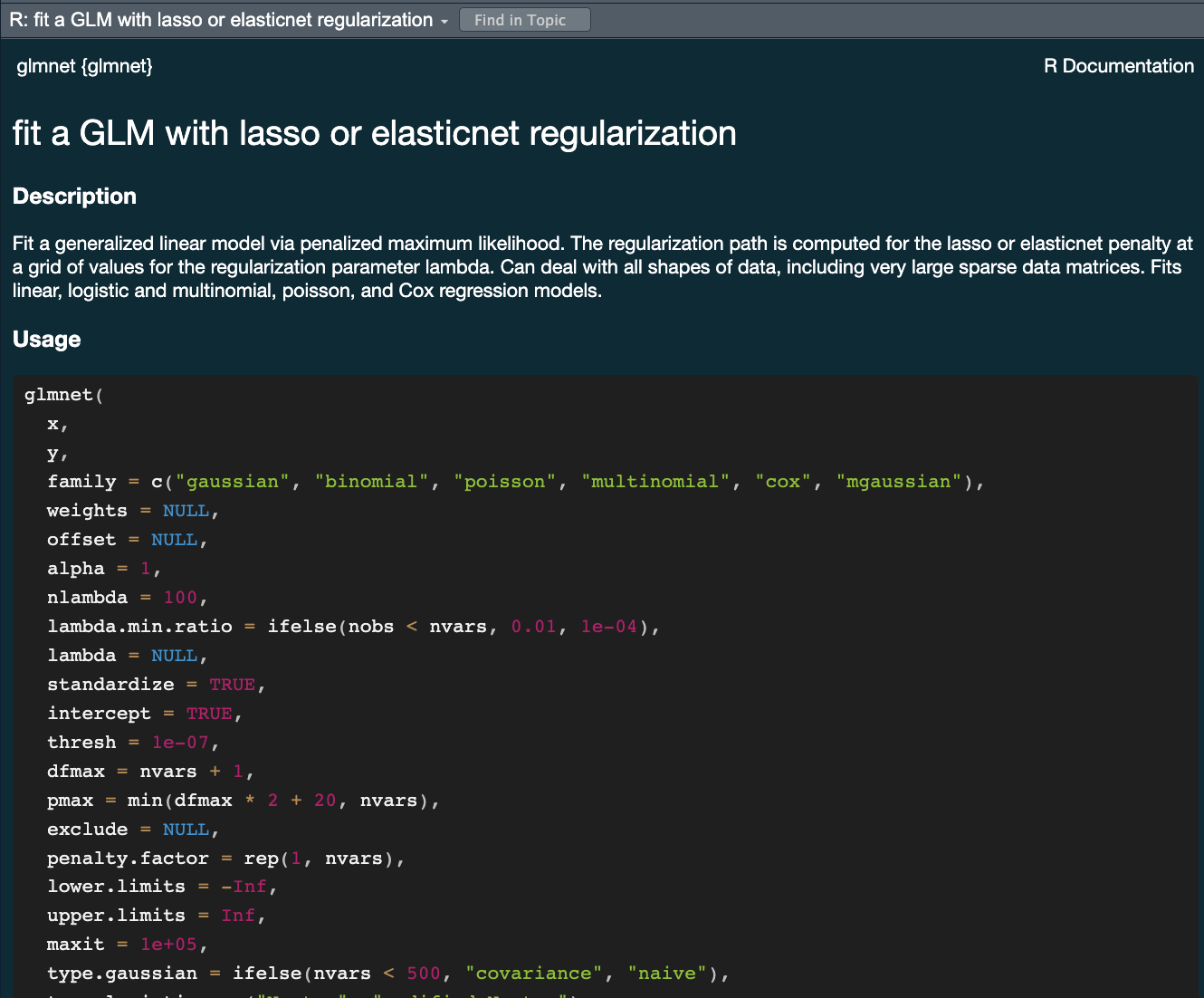
The tidymodels advantage
- In the
tidymodelsframework, all models are created using the same syntax:- This makes it easy to compare models
- This makes it easy to switch between models
- This makes it easy to use the same model with different engines (packages)
- The
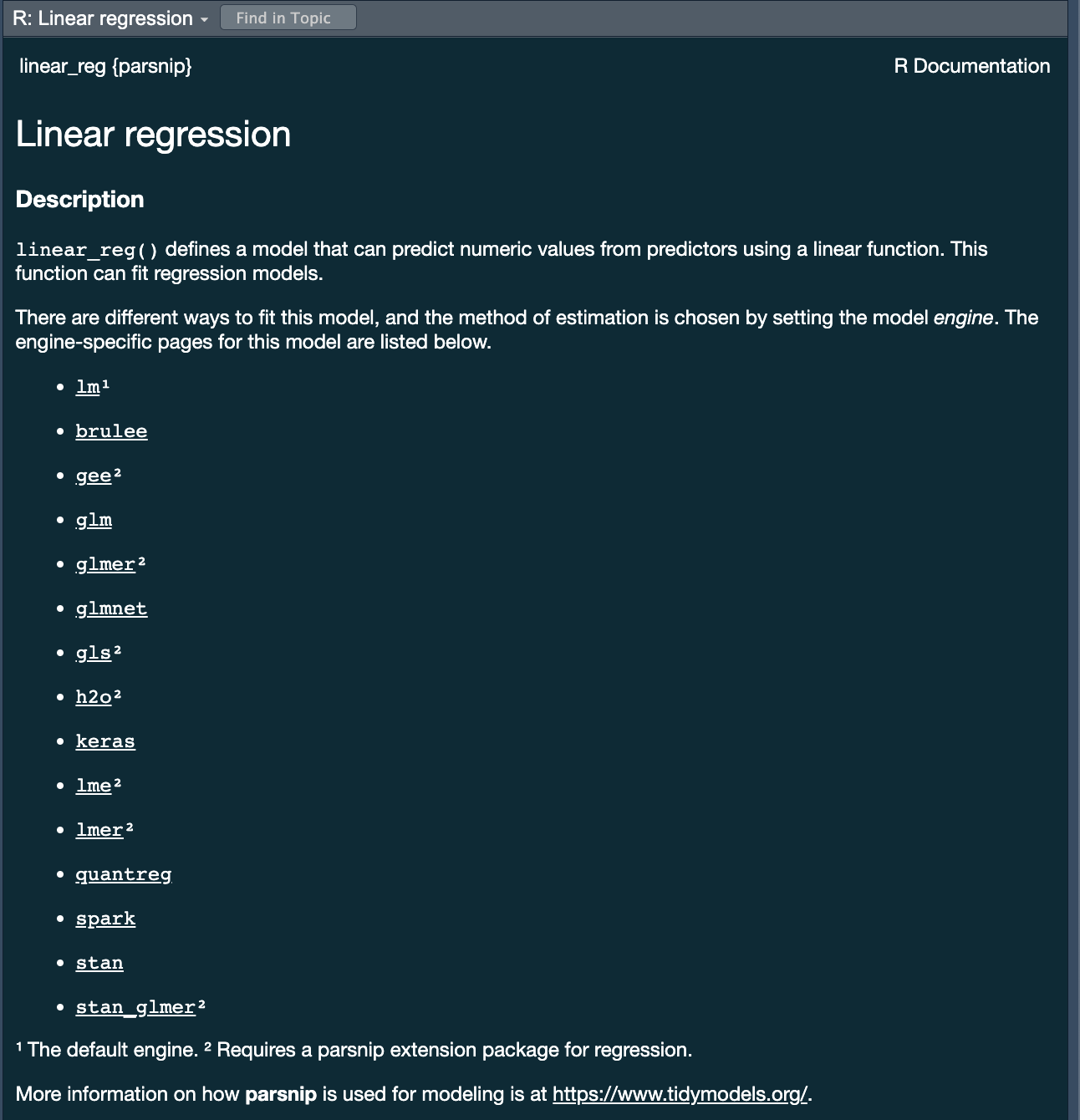
parsnippackage provides a consistent interface to many models - For example, to fit a linear model you would be able to access the
linear_reg()function
A tidymodels prediction will … ![]()
- always be inside a tibble
- have column names and types that are unsurprising and predictable
- ensure the number of rows in
new_dataand the output are the same
To specify a model ![]()
- Choose a model
- Specify an engine
- Set the mode
Specify a model: Type ![]()
Next week we will discuss model types more thoroughly. For now, we will focus on two types:
- Logistic Regression
We chose these two because they are robust, simple models that fit our goal of predicting a binary condition/class (forested/not forested)
Specify a model: engine ![]()
Specify a model: engine ![]()
Specify a model: mode ![]()
Specify a model: mode ![]()
Some models have a limited set of modes …
. . .
Others require specification …
All available models are listed at https://www.tidymodels.org/find/parsnip/
To specify a model ![]()
- Choose a model
- Specify an engine
- Set the mode
Two algorithms to try
We have a problem (predict whether a hexagon is forested), data (forested_train), and a way to evaluate predictions. Before we wire up the full tidymodels plumbing, let’s look at the two models we’ll compare today.
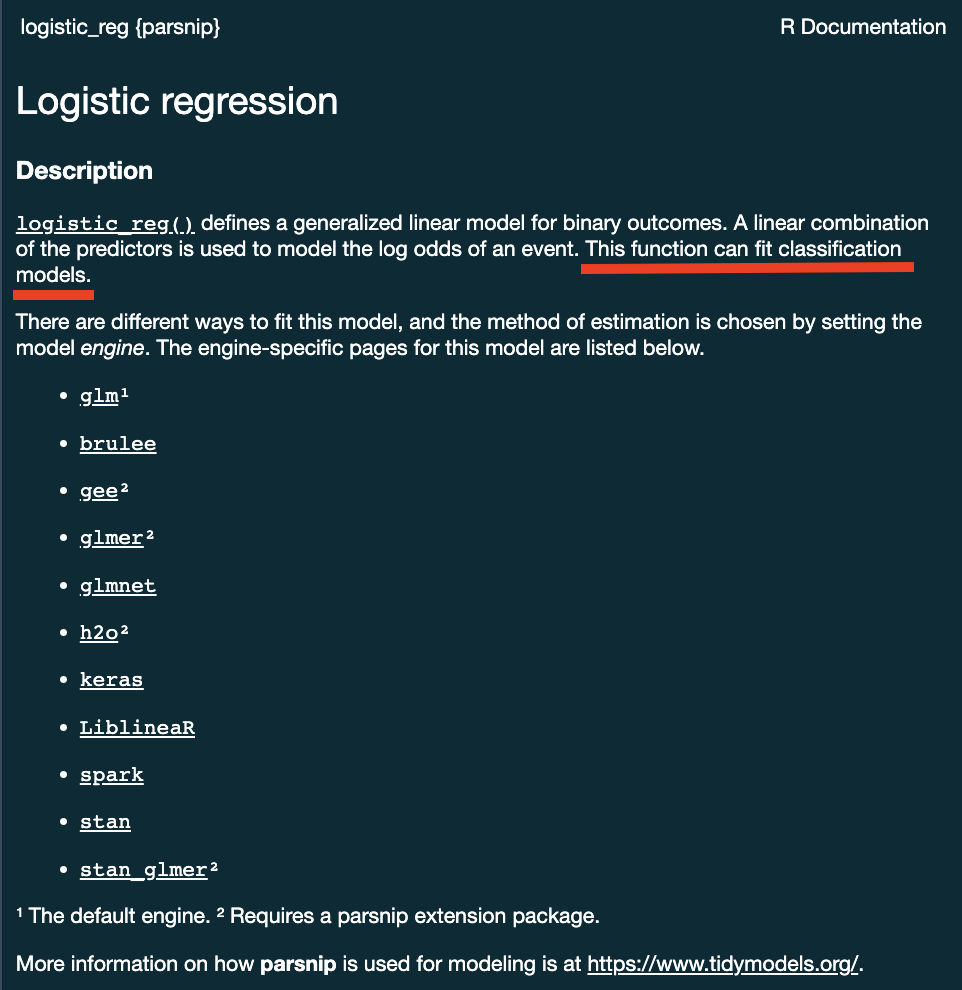
Logistic regression
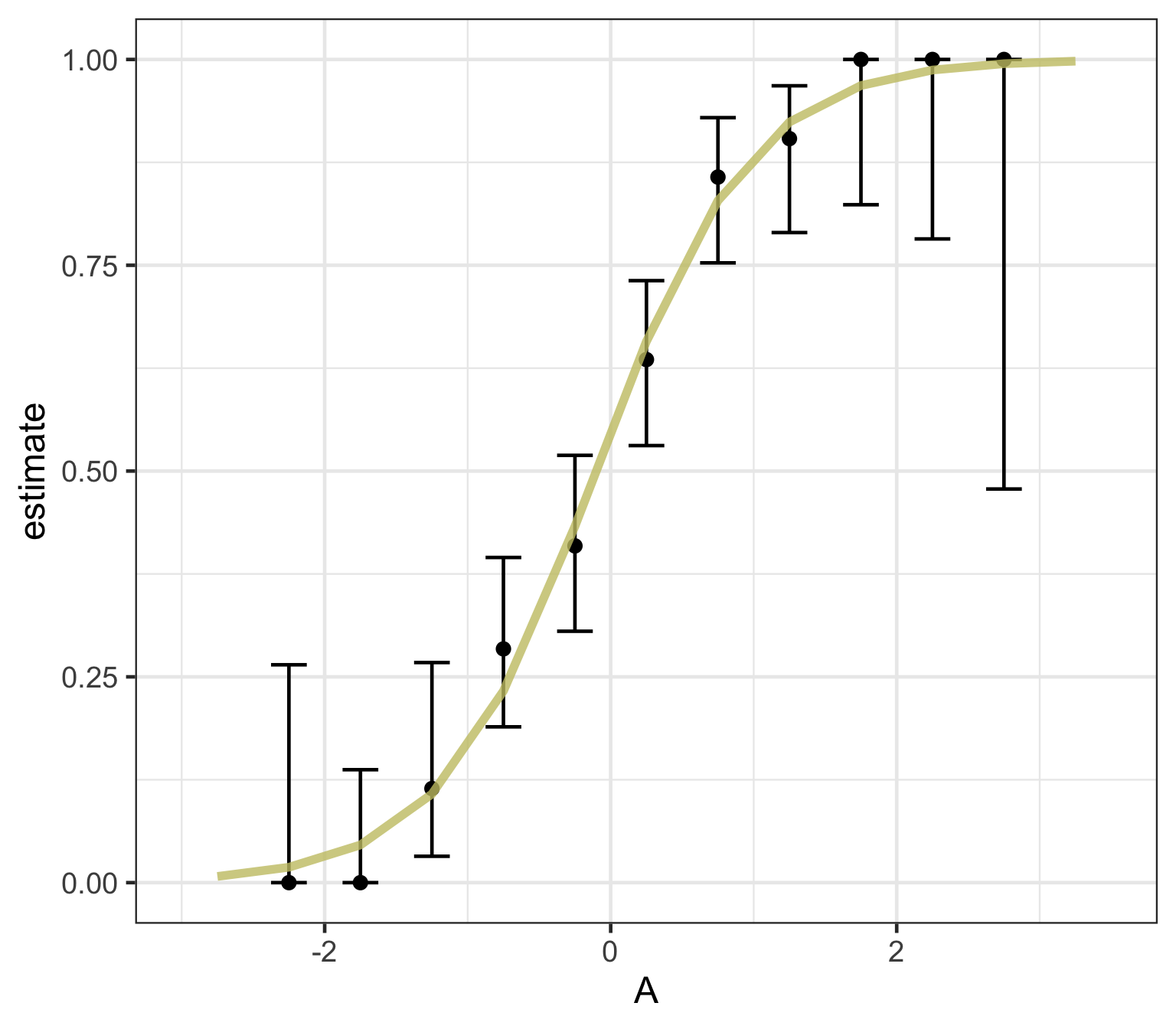
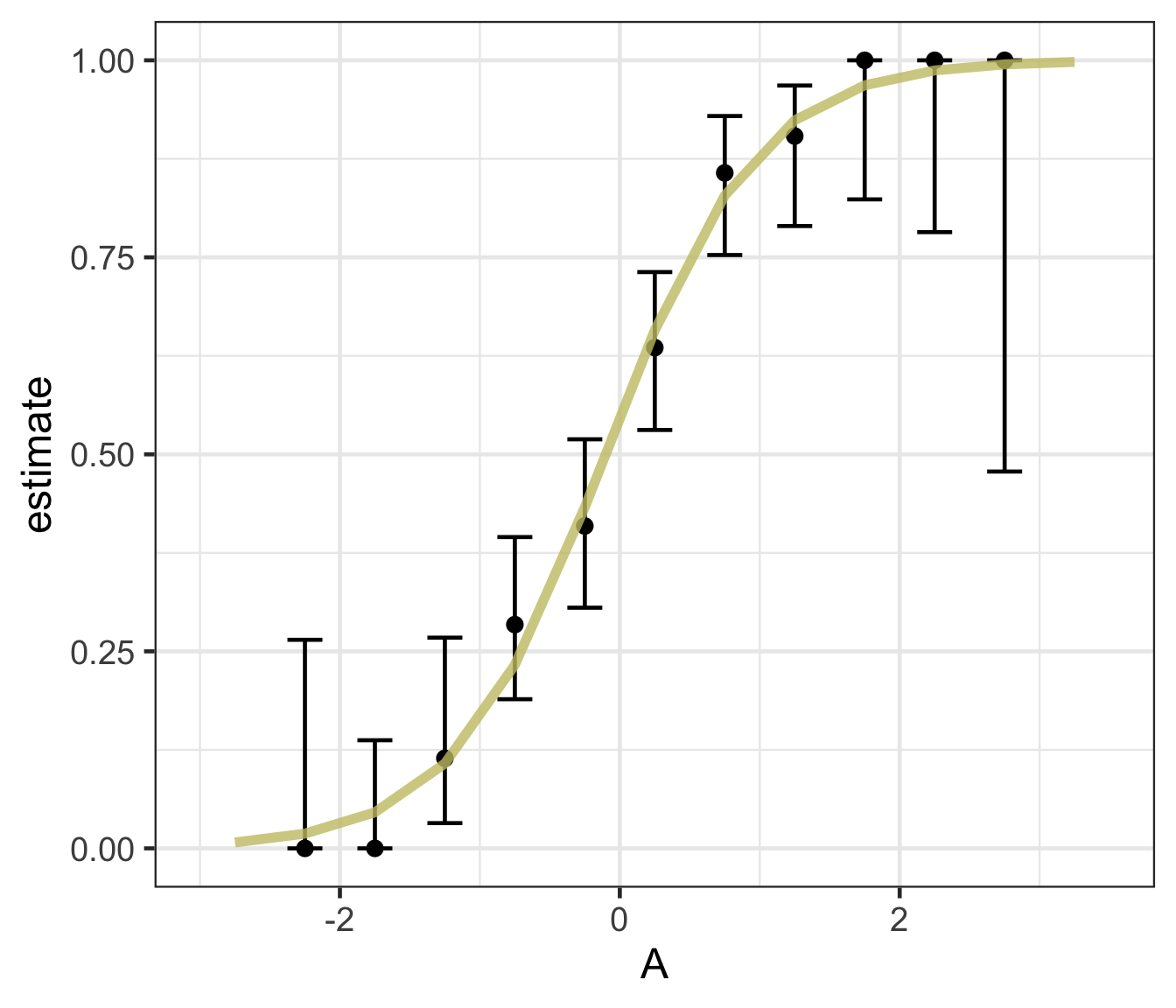
Logistic regression predicts probability—instead of a straight line, it gives an S-shaped curve that estimates how likely an outcome (e.g., is forested) is based on a predictor (e.g., rainfall and temperature).
The dots in the plot show the actual proportion of “successes” (e.g., is forested) within different bins of the predictor variable(s) (A).
The vertical error bars represent uncertainty—showing a range where the true probability might fall.
Logistic regression helps answer “how likely” questions—e.g., “How likely is someone to pass based on their study hours?” rather than just predicting a yes/no outcome.
Decision trees — structure & pruning
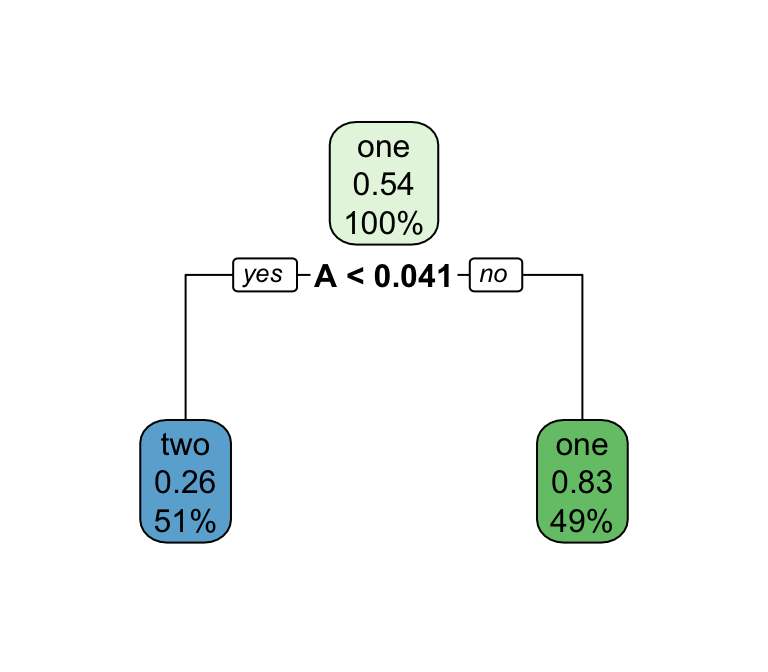
Series of splits or if/then statements based on predictors
First the tree grows until some condition is met (maximum depth, no more data)
Then the tree is pruned to reduce its complexity
Each terminal leaf predicts a class (or, for regression, a numeric value)
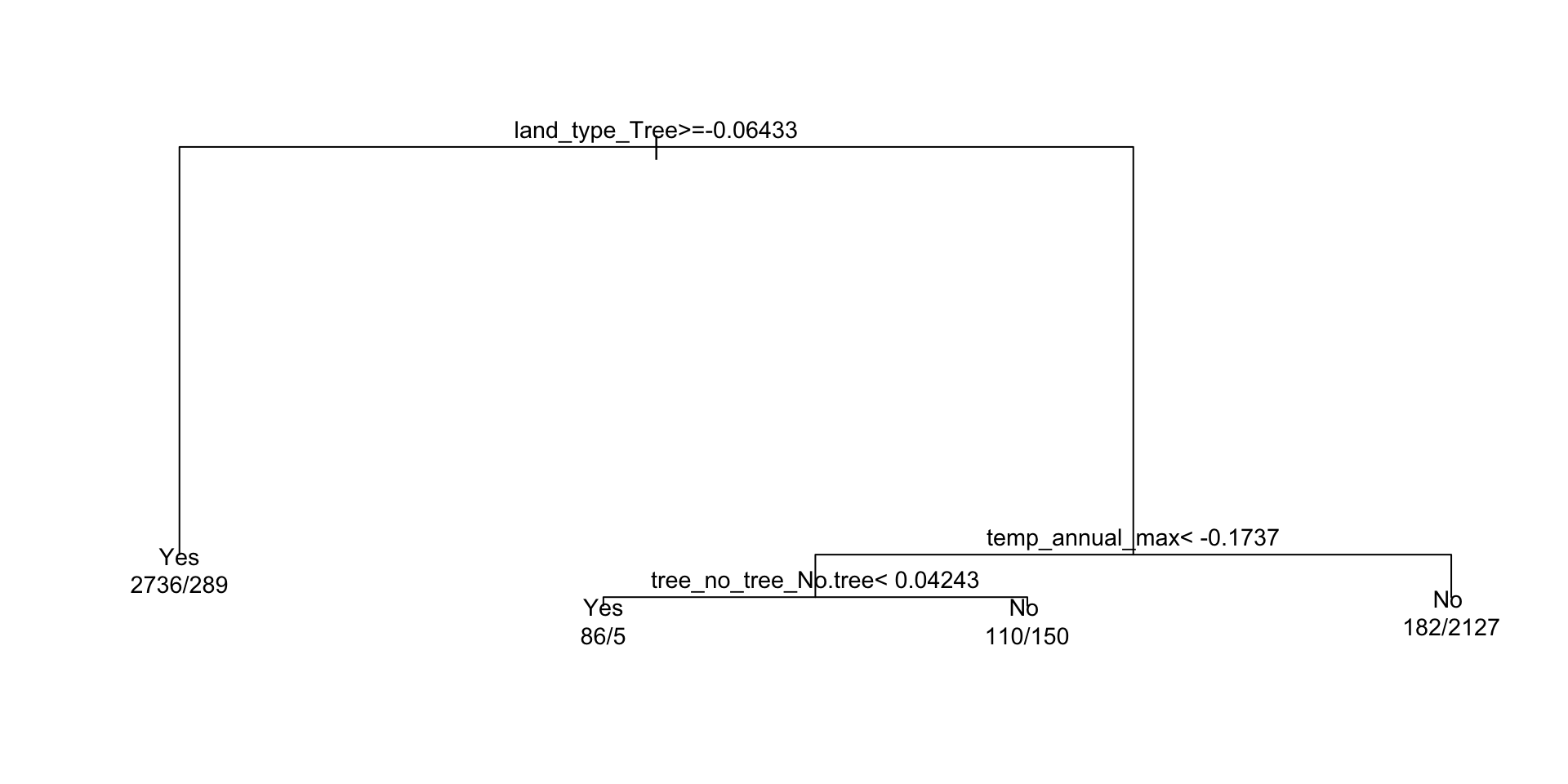
Decision trees — what predictions look like
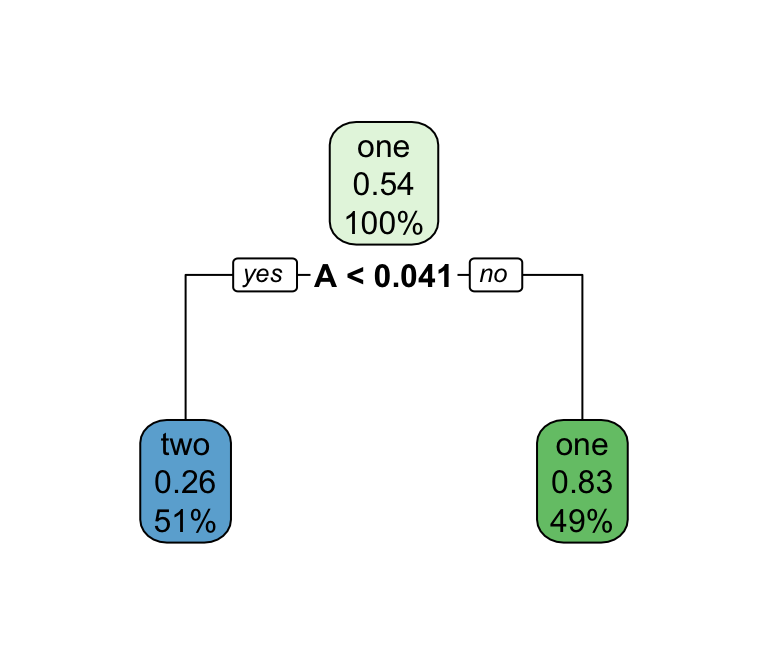
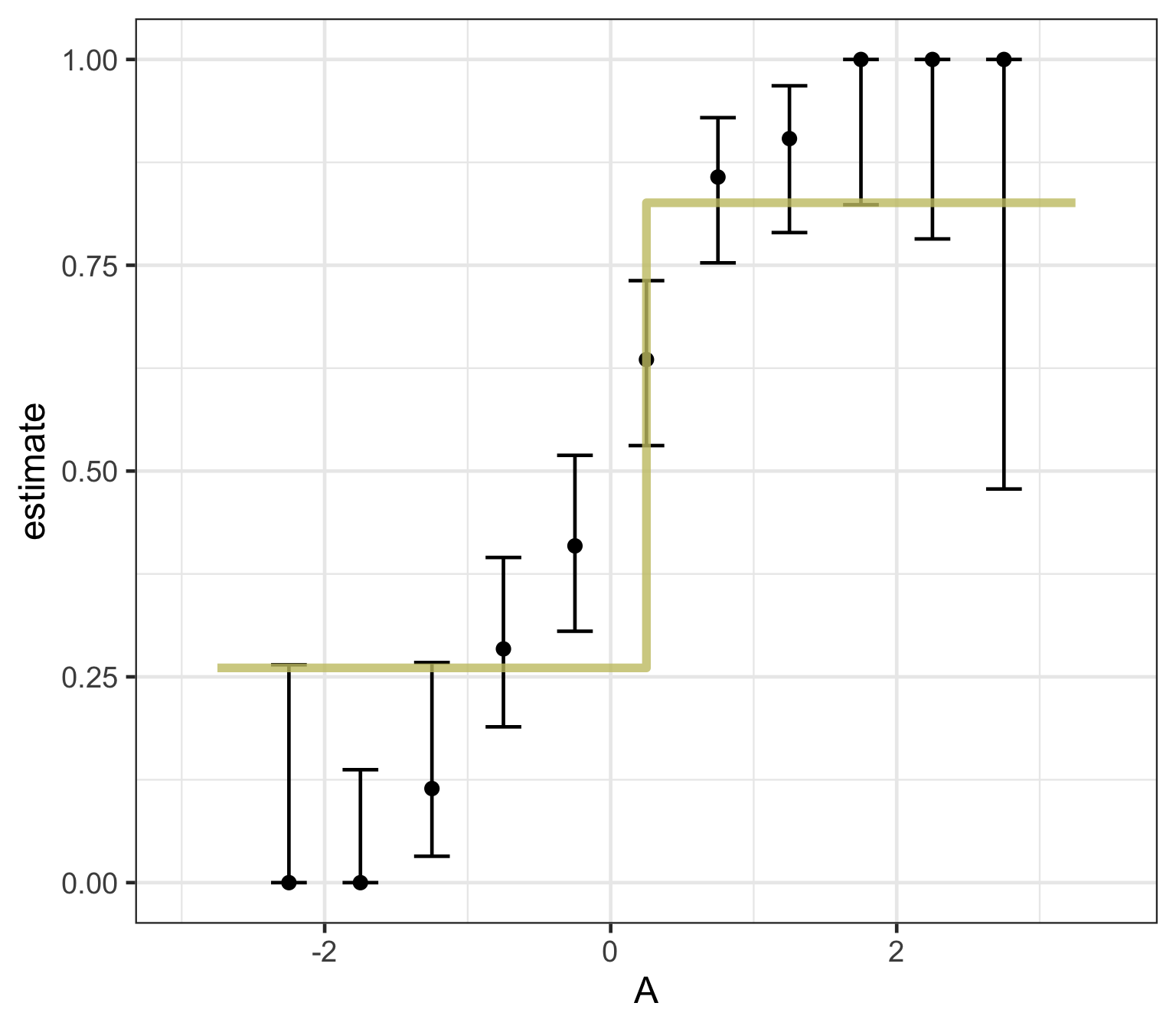
A tree’s decision boundary is stepwise — in contrast to the smooth S-curve of logistic regression.
When would you reach for each?
Logistic regression
- Smooth, monotonic decision boundary
- Returns calibrated probabilities, not just a class
- Coefficients are interpretable (log-odds per predictor)
- Sensitive to predictor scale → benefits from normalization
- Assumes a (largely) linear relationship on the log-odds scale
Decision trees
- Step-wise, rectangular splits on predictors
- Handles non-linearities and interactions natively
- No need to scale, dummy-encode, or worry about collinearity
- Easy to explain visually, but a single tree can overfit and is unstable across samples
- Sets up ensemble methods (random forest, boosting) next week
What algorithm is best for estimating forested plots?
We can only really know by testing them … but we know the options!
Logistic regression
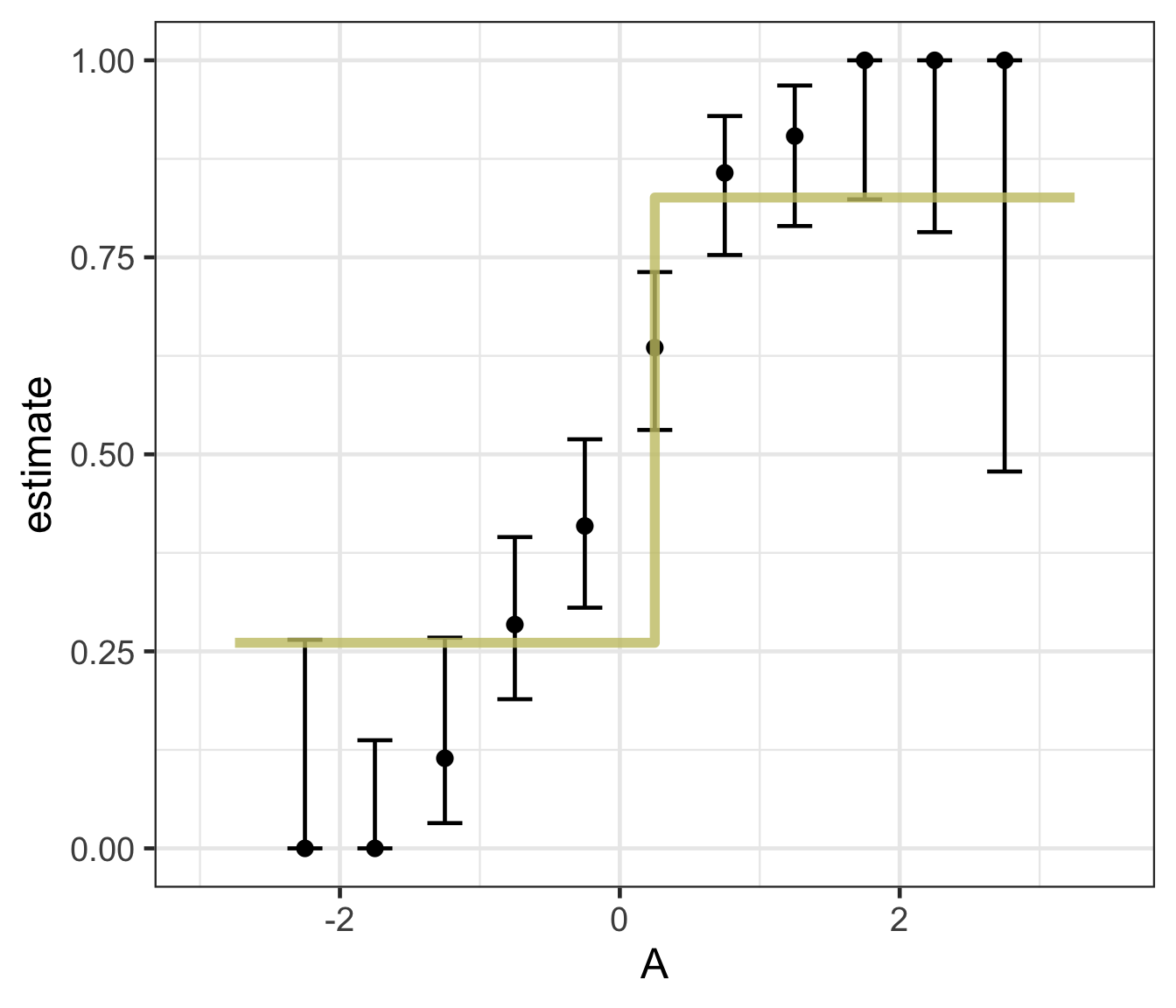
Decision trees
Feature engineering with recipes
Why preprocess? ![]()
A raw formula (
forested ~ .) would send features to the model untouched.But many models care how their features look — different scales, skewed distributions, missing values, or categorical levels can bias the fit.
recipeslets us declare a preprocessing pipeline once, estimate its parameters from the training data, and apply it consistently to every new data set (folds, test set, future predictions).
Important
Preprocessing parameters (means, sds, imputed values, PCA loadings) are learned only on the training set — this prevents data leakage from test data into the fit.
Data Normalization in tidymodels
- We’ve now seen why distributions matter and how to detect non-normality.
- In the
recipesframework, normalization is declared once and applied consistently to every split — preventing data leakage. - The same
step_*()functions handle min-max scaling, z-score standardization, power transforms, and more.
Common Normalization Techniques
| Technique | Idea | When to use | step_* |
|---|---|---|---|
| Min-max | rescale to \([0, 1]\): \(x' = \frac{x - x_\text{min}}{x_\text{max} - x_\text{min}}\) | Need a bounded range; beware outliers | step_range() |
| Z-score | center + scale: \(x' = \frac{x - \mu}{\sigma}\) | Symmetric-ish features, different units | step_normalize() |
| Power | Box-Cox (positive) or Yeo-Johnson (any sign) estimate an optimal power transform | Reshape skew toward normality before scaling | step_YeoJohnson(), step_BoxCox() |
| Log | \(x' = \log(x)\) | Right-skewed positive data (e.g. streamflow) | step_log() |
| PCA | project to a lower-dimensional basis that preserves variance | Many correlated numeric predictors | step_pca() |
Tip
For environmental variables (streamflow, precipitation, concentrations) log is by far the most common first move — they’re almost always right-skewed.
Other Feature Engineering Tasks
Beyond scaling, three recipes moves you’ll reach for most often in water resources:
Handling missing values
step_impute_mean()/step_impute_median()— fast, parametric fillstep_impute_knn()— use neighboring rows (preserves covariance structure)
Encoding categorical variables
step_dummy(all_nominal_predictors())— one-hot encodes factorsstep_other()— collapses rare levels into “other”
Creating interaction terms
step_interact(terms = ~ aridity:p_mean)— add variable products when the effect of one predictor depends on another
Note
The full step_* catalog (text, splines, binning, dimension reduction, …) lives at https://recipes.tidymodels.org/reference/. Search there when you need something not listed here.
Implementing a recipe in tidymodels
Declare a
recipe()with a formula + training data.Add
step_*()preprocessing operations (imputation, dummies, normalization, …).prep()estimates the transformation parameters from the training data.bake()applies those transformations to any data set (training, test, new).
Example Workflow
library(tidymodels)
(recipe_obj <- recipe(flipper_length_mm ~
bill_length_mm + body_mass_g +
sex + island + species,
data = penguins) |>
step_impute_mean(all_numeric_predictors()) |>
step_dummy(all_factor_predictors()) |>
step_normalize(all_numeric_predictors()))
# Prepare and apply transformations
prep_recipe <- prep(recipe_obj, training = penguins)
normalized_data <- bake(prep_recipe, new_data = NULL) |>
mutate(species = penguins$species) |>
drop_na()
head(normalized_data)
#> # A tibble: 6 × 9
#> bill_length_mm body_mass_g flipper_length_mm sex_male island_Dream
#> <dbl> <dbl> <int> <dbl> <dbl>
#> 1 -0.895 -0.568 181 0.990 -0.764
#> 2 -0.822 -0.506 186 -1.01 -0.764
#> 3 -0.675 -1.19 195 -1.01 -0.764
#> 4 -1.33 -0.940 193 -1.01 -0.764
#> 5 -0.858 -0.692 190 0.990 -0.764
#> 6 -0.931 -0.723 181 -1.01 -0.764
#> # ℹ 4 more variables: island_Torgersen <dbl>, species_Chinstrap <dbl>,
#> # species_Gentoo <dbl>, species <fct>Explanation:
recipe()defines preprocessing steps for modeling.step_impute_mean(all_numeric_predictors())fills missing numeric values with the column mean (estimated on training data).step_dummy(all_factor_predictors())creates dummy variables for categorical predictors.step_normalize(all_numeric_predictors())standardizes all numerical predictors (mean 0, sd 1).prep()estimates parameters for transformations.bake()applies the transformations to the dataset.
A recipe for our running example ![]()
Back to forested. Before we fit any models, let’s declare the preprocessing pipeline we want every model in this deck to use — impute missing numeric values, dummy-encode the nominal predictors, and normalize everything numeric:
Note
We’ll reuse forested_recipe for every model we specify for the rest of today (decision tree, logistic regression, workflow set) — the point of a recipe is to define preprocessing once and reuse it everywhere. The same recipe carries forward into Wednesday’s lecture (VIP + spatial CV) and next week’s (more model families + tuning).
Fitting your model:
- OK! So you have split data, a recipe, and a model spec (with engine + mode)
- Now you need to fit the model!
- Before we reach for
workflow(), it’s useful to see how a recipe + model compose manually:prep()the recipe on the training data,bake()it to get the transformed frame, then hand that frame tofit().
dt_model <- decision_tree() %>%
set_engine('rpart') %>%
set_mode("classification")
# Estimate preprocessing parameters (means, dummy levels, etc.) from training data
prepped <- prep(forested_recipe, training = forested_train)
# Apply the pipeline
forested_baked <- bake(prepped, new_data = NULL)
# Fit the model against the fully preprocessed training data
# Note: `.` now picks up the dummy-encoded + normalized columns produced by the recipe
(output <- fit(dt_model, forested ~ ., data = forested_baked))
#> parsnip model object
#>
#> n= 5685
#>
#> node), split, n, loss, yval, (yprob)
#> * denotes terminal node
#>
#> 1) root 5685 2571 Yes (0.54775726 0.45224274)
#> 2) land_type_Tree>=-0.06433113 3025 289 Yes (0.90446281 0.09553719) *
#> 3) land_type_Tree< -0.06433113 2660 378 No (0.14210526 0.85789474)
#> 6) temp_annual_max< -0.1737334 351 155 Yes (0.55840456 0.44159544)
#> 12) tree_no_tree_No.tree< 0.04242667 91 5 Yes (0.94505495 0.05494505) *
#> 13) tree_no_tree_No.tree>=0.04242667 260 110 No (0.42307692 0.57692308) *
#> 7) temp_annual_max>=-0.1737334 2309 182 No (0.07882200 0.92117800) *Tip
This prep → bake → fit dance is what workflow() automates on the next slide. Seeing it once makes the workflow version less magical.
Component extracts:
While
tidymodelsprovides a standard interface to all models all base models have specific methods (remembersummary.lm? )To use these, you must extract the underlying object.
extract_fit_engine()extracts the underlying model objectextract_fit_parsnip()extracts the parsnip objectextract_recipe()extracts the recipe objectextract_preprocessor()extracts the preprocessor object… many more
A model workflow
Workflows — a container for your pipeline ![]()
The workflows package bundles a preprocessor (recipe or formula) and a model spec into a single object. Instead of managing prep → bake → fit by hand, you hand the workflow your training data and it does the bookkeeping on every resample.
The four-step pattern:
- Create a workflow (
workflow()— likeggplot()instantiates a canvas) - Add a preprocessor (
add_recipe()oradd_formula()) - Add a model (
add_model()) - Fit it (
fit()orfit_resamples())
Why bother?
✅ Preprocessing and modeling travel together — no forgetting to bake the test set ✅ Same skeleton for every model — swap in a new add_model() spec and go ✅ Works identically with a single fit(), resamples, or tuning
A model workflow ![]()
![]()
Instead of prep/bake/fit by hand, drop the recipe and the model into a workflow() and let it do the bookkeeping:
#> ══ Workflow [trained] ══════════════════════════════════════════════════════════
#> Preprocessor: Recipe
#> Model: decision_tree()
#>
#> ── Preprocessor ────────────────────────────────────────────────────────────────
#> 3 Recipe Steps
#>
#> • step_impute_mean()
#> • step_dummy()
#> • step_normalize()
#>
#> ── Model ───────────────────────────────────────────────────────────────────────
#> n= 5685
#>
#> node), split, n, loss, yval, (yprob)
#> * denotes terminal node
#>
#> 1) root 5685 2571 Yes (0.54775726 0.45224274)
#> 2) land_type_Tree>=-0.06433113 3025 289 Yes (0.90446281 0.09553719) *
#> 3) land_type_Tree< -0.06433113 2660 378 No (0.14210526 0.85789474)
#> 6) temp_annual_max< -0.1737334 351 155 Yes (0.55840456 0.44159544)
#> 12) tree_no_tree_No.tree< 0.04242667 91 5 Yes (0.94505495 0.05494505) *
#> 13) tree_no_tree_No.tree>=0.04242667 260 110 No (0.42307692 0.57692308) *
#> 7) temp_annual_max>=-0.1737334 2309 182 No (0.07882200 0.92117800) *Tip
add_recipe() means the workflow carries your feature engineering pipeline and the model together — fit() automatically prep()/bake()s for every resample. (You can use add_formula() instead if you don’t need any preprocessing, but once you have a recipe there’s no reason to.)
Predict with your model ![]()
![]()
How do you apply your fitted workflow to the test set?
augment()on a workflow acceptsnew_data =and returns the test data with predictions appended.
#> # A tibble: 1,422 × 22
#> .pred_class .pred_Yes .pred_No forested year elevation eastness northness
#> <fct> <dbl> <dbl> <fct> <dbl> <dbl> <dbl> <dbl>
#> 1 Yes 0.904 0.0955 No 2005 164 -84 53
#> 2 Yes 0.904 0.0955 Yes 2005 806 47 -88
#> 3 No 0.423 0.577 Yes 2005 2240 -67 -74
#> 4 Yes 0.904 0.0955 Yes 2005 787 66 -74
#> 5 Yes 0.904 0.0955 Yes 2005 1330 99 7
#> 6 Yes 0.904 0.0955 Yes 2005 1423 46 88
#> 7 Yes 0.904 0.0955 Yes 2014 546 -92 -38
#> 8 Yes 0.904 0.0955 Yes 2014 1612 30 -95
#> 9 No 0.0788 0.921 No 2014 235 0 -100
#> 10 No 0.0788 0.921 No 2014 308 -70 -70
#> # ℹ 1,412 more rows
#> # ℹ 14 more variables: roughness <dbl>, tree_no_tree <fct>, dew_temp <dbl>,
#> # precip_annual <dbl>, temp_annual_mean <dbl>, temp_annual_min <dbl>,
#> # temp_annual_max <dbl>, temp_january_min <dbl>, vapor_min <dbl>,
#> # vapor_max <dbl>, canopy_cover <dbl>, lon <dbl>, lat <dbl>, land_type <fct>Evaluate your model ![]()
How do you evaluate the skill of your models?
We will learn more about model evaluation and tuning later in this unit, but for now we can blindly use the metrics() function defaults to get a sense of how our model is doing.
The default metrics for classification models are accuracy and kap (Cohen’s Kappa)
The default metrics for regression models are rmse, rsq (R²), and mae
tidymodels advantage! ![]()
- OK! while that was not too much work, it certainly wasn’t minimal.
Now let’s say you want to check if the logistic regression model performs better than the decision tree.
That sounds like a lot to repeat.
Fortunately, the
tidymodelsframework makes this a straightforward swap!
log_mod = logistic_reg() %>%
set_engine("glm") %>%
set_mode("classification")
log_wf <- workflow() %>%
add_recipe(forested_recipe) %>%
add_model(log_mod) %>%
fit(data = forested_train)
log_preds <- augment(log_wf, new_data = forested_test)
metrics(log_preds, truth = forested, estimate = .pred_class)
#> # A tibble: 2 × 3
#> .metric .estimator .estimate
#> <chr> <chr> <dbl>
#> 1 accuracy binary 0.890
#> 2 kap binary 0.777Note
Note how little changed: same recipe, same workflow skeleton — only the model spec is different. That swap is the whole point of tidymodels.
So who wins?
metrics(dt_preds, truth = forested, estimate = .pred_class)
#> # A tibble: 2 × 3
#> .metric .estimator .estimate
#> <chr> <chr> <dbl>
#> 1 accuracy binary 0.885
#> 2 kap binary 0.768
metrics(log_preds, truth = forested, estimate = .pred_class)
#> # A tibble: 2 × 3
#> .metric .estimator .estimate
#> <chr> <chr> <dbl>
#> 1 accuracy binary 0.890
#> 2 kap binary 0.777Right out of the gate, there doesn’t seem to be much difference, but …
Don’t get too confident — use resamples
- We have a single point-estimate of performance — one train/test split produces one number that can be misleadingly high or low depending on which rows ended up in the test set.
- Resampling repeatedly holds out different subsets, fits the model each time, and averages the metric — giving a more stable and trustworthy estimate of skill.
- Instead, we can use our
resamplesto better understand the skill of a model based on the iterative leave-out policy:
#> # A tibble: 1 × 5
#> splits id .metrics .notes .predictions
#> <list> <chr> <list> <list> <list>
#> 1 <split [5116/569]> Fold01 <tibble [3 × 4]> <tibble [0 × 4]> <tibble [569 × 6]>Note
tidyr::unnest could be used to expand any of those list columns!
Don’t get too confident — same pattern, other model
- We can execute the same workflow for the logistic regression model, this time evaluating the resamples instead of the test data.
#> # Resampling results
#> # 10-fold cross-validation
#> # A tibble: 10 × 5
#> splits id .metrics .notes .predictions
#> <list> <chr> <list> <list> <list>
#> 1 <split [5116/569]> Fold01 <tibble [3 × 4]> <tibble [0 × 4]> <tibble>
#> 2 <split [5116/569]> Fold02 <tibble [3 × 4]> <tibble [0 × 4]> <tibble>
#> 3 <split [5116/569]> Fold03 <tibble [3 × 4]> <tibble [0 × 4]> <tibble>
#> 4 <split [5116/569]> Fold04 <tibble [3 × 4]> <tibble [0 × 4]> <tibble>
#> 5 <split [5116/569]> Fold05 <tibble [3 × 4]> <tibble [0 × 4]> <tibble>
#> 6 <split [5117/568]> Fold06 <tibble [3 × 4]> <tibble [0 × 4]> <tibble>
#> 7 <split [5117/568]> Fold07 <tibble [3 × 4]> <tibble [0 × 4]> <tibble>
#> 8 <split [5117/568]> Fold08 <tibble [3 × 4]> <tibble [0 × 4]> <tibble>
#> 9 <split [5117/568]> Fold09 <tibble [3 × 4]> <tibble [0 × 4]> <tibble>
#> 10 <split [5117/568]> Fold10 <tibble [3 × 4]> <tibble [0 × 4]> <tibble>Don’t get too confident — collect the metrics
Since we now have an “ensemble” of models (across the folds), we can collect a summary of the metrics across them.
The default metrics for classification ensembles are:
- accuracy: accuracy is the proportion of correct predictions
- brier_class: the Brier score for classification is the mean squared difference between the predicted probability and the actual outcome
- roc_auc: the area under the ROC (Receiver Operating Characteristic) curve
collect_metrics(log_wf_rs)
#> # A tibble: 3 × 6
#> .metric .estimator mean n std_err .config
#> <chr> <chr> <dbl> <int> <dbl> <chr>
#> 1 accuracy binary 0.906 10 0.00360 pre0_mod0_post0
#> 2 brier_class binary 0.0714 10 0.00191 pre0_mod0_post0
#> 3 roc_auc binary 0.961 10 0.00164 pre0_mod0_post0
collect_metrics(dt_wf_rs)
#> # A tibble: 3 × 6
#> .metric .estimator mean n std_err .config
#> <chr> <chr> <dbl> <int> <dbl> <chr>
#> 1 accuracy binary 0.896 10 0.00385 pre0_mod0_post0
#> 2 brier_class binary 0.0889 10 0.00224 pre0_mod0_post0
#> 3 roc_auc binary 0.909 10 0.00267 pre0_mod0_post0Further simplification: workflowsets
OK, so we have streamlined a lot of things with tidymodels:
We have a specified preprocessor (e.g. formula or recipe)
We have defined a model (complete with engine and mode)
We have implemented workflows pairing each model with the preprocessor to either fit the model to the resamples, or, the training data.
While a significant improvement, we can do better!
Note
Does the idea of implementing a process over a list of elements ring a bell?
Model mappings: ![]()
workflowsetsis a package that builds off ofpurrrand allows you to iterate over multiple models and/or multiple resamplesRemember how
map,map_*,map2, andwalk2functions allow lists to map to lists or vectors?workflow_setmaps a preprocessor (formula or recipe) to a set of models - each provided as alistobject.To start, we will create a
workflow_setobject (instead of aworkflow). The first argument is a list of preprocessor objects (formulas or recipes) and the second argument is a list of model objects.Both must be lists, by nature of the underlying
purrrcode.
Iterative execution
Once the
workflow_setobject is created, theworkflow_mapfunction can be used to map a function across the preprocessor/model combinations.Here we are mapping the
fit_resamplesfunction across theworkflow_setcombinations using theresamplesargument to specify the resamples we want to use (forested_folds).
#> # A workflow set/tibble: 2 × 4
#> wflow_id info option result
#> <chr> <list> <list> <list>
#> 1 recipe_logistic_reg <tibble [1 × 4]> <opts[1]> <rsmp[+]>
#> 2 recipe_decision_tree <tibble [1 × 4]> <opts[1]> <rsmp[+]>Rapid comparison
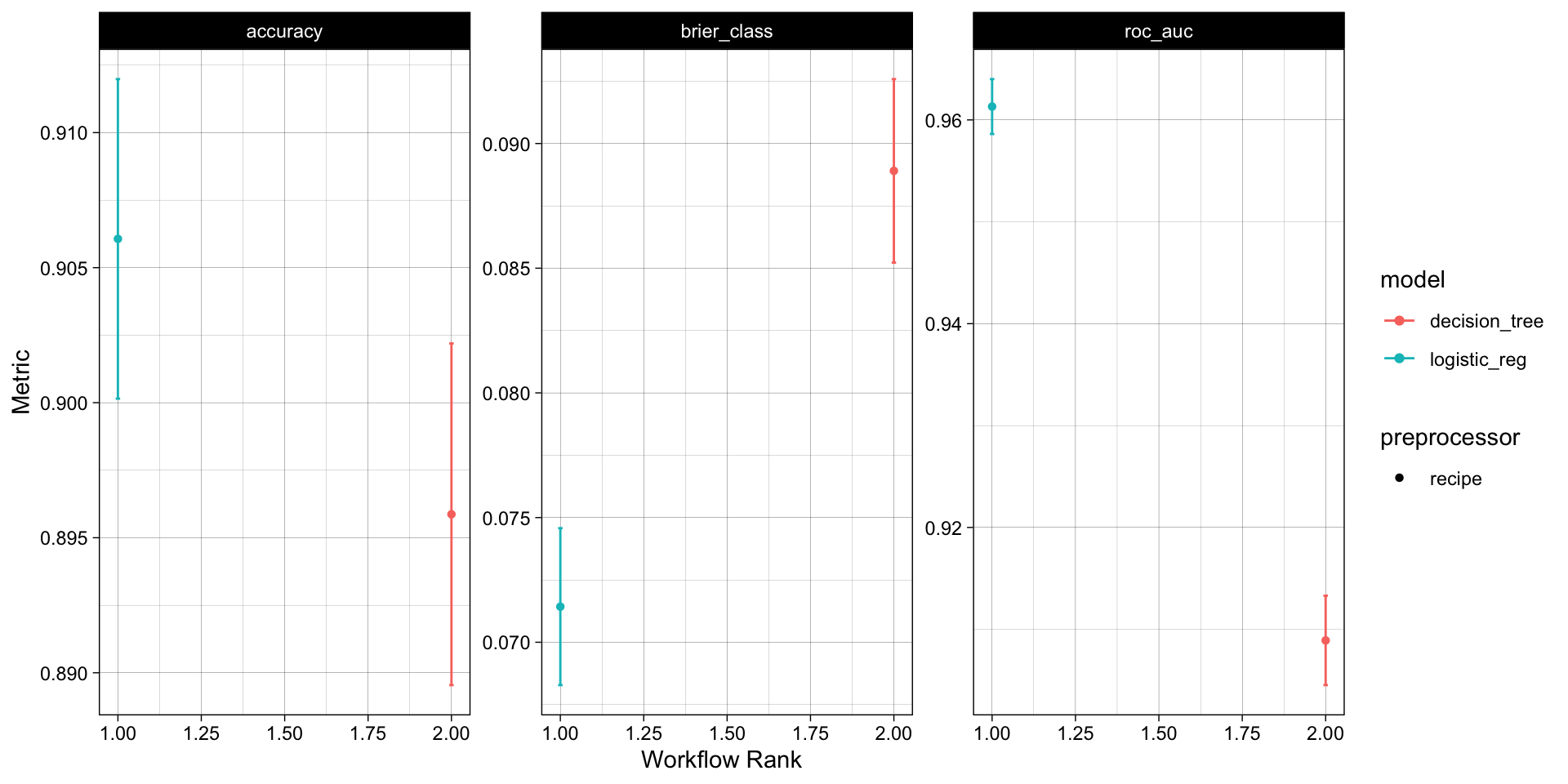
With that single function, all models have been fit to the resamples and we can quickly compare the results both graphically and statistically:
# Long list of results, ranked by accuracy
rank_results(wf_obj,
rank_metric = "accuracy",
select_best = TRUE)
#> # A tibble: 6 × 9
#> wflow_id .config .metric mean std_err n preprocessor model rank
#> <chr> <chr> <chr> <dbl> <dbl> <int> <chr> <chr> <int>
#> 1 recipe_logistic… pre0_m… accura… 0.906 0.00360 10 recipe logi… 1
#> 2 recipe_logistic… pre0_m… brier_… 0.0714 0.00191 10 recipe logi… 1
#> 3 recipe_logistic… pre0_m… roc_auc 0.961 0.00164 10 recipe logi… 1
#> 4 recipe_decision… pre0_m… accura… 0.896 0.00385 10 recipe deci… 2
#> 5 recipe_decision… pre0_m… brier_… 0.0889 0.00224 10 recipe deci… 2
#> 6 recipe_decision… pre0_m… roc_auc 0.909 0.00267 10 recipe deci… 2Overall, the logistic regression model appears to be the best model for this data set!
Note
Keep in mind: we compared these models out of the box, with no feature engineering, no hyperparameter tuning, and a single resampling scheme. Next week we’ll revisit this comparison once we can actually tune a model 👀
The whole game - status update
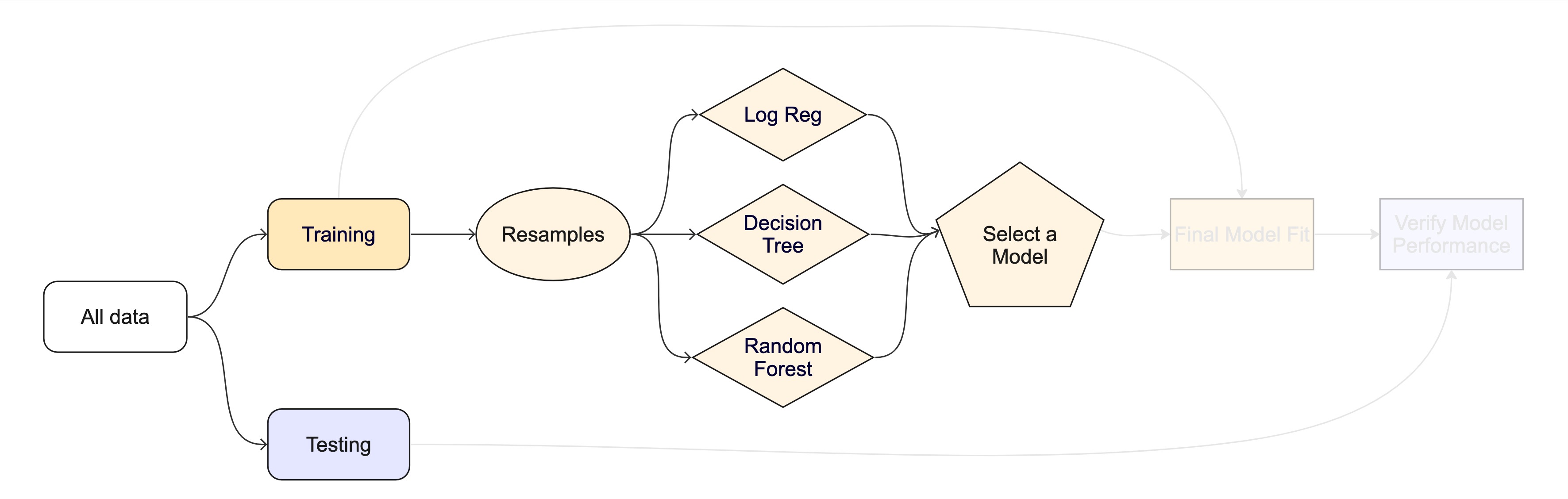
On Wednesday we’ll ask two things collect_metrics() can’t answer: which predictors did the model actually use (VIP) and is our 0.9-ish score honest (spatial cross-validation on autocorrelated data). Next week we’ll meet a bigger zoo of model families and learn to tune them.