Week 5
Beyond collect_metrics(): Interpretation & Spatial Resampling
Where we left off
On Monday we built a full tidymodels pipeline on the forested dataset:
initial_split()→training()/testing()vfold_cv()→ 10-fold cross-validation on the training setrecipe()→ impute → dummy → normalizeparsnipspec →workflow()→fit_resamples()collect_metrics()→ accuracy, ROC AUC, Brier score
A logistic regression came out ahead of a decision tree, and we stopped there.
Today: two questions collect_metrics() can’t answer
“Which predictors actually drove the prediction?” — a global ROC AUC doesn’t tell you whether the model leaned on
elevation,canopy_cover, orvapor_max.“Is my 0.92 AUC honest?” — or did the model just memorize spatial context, because neighboring hexagons in Washington state are nearly identical?
Today’s two tools:
- 📊
vip— Variable Importance Plots: rank predictors by how much a model relied on them. - 🗺️
spatialsample— Spatial resampling: cross-validation schemes that respect the geography of your data.
And a key insight: the first tool is not trustworthy without the second.
By the end of today you will be able to…
- Extract a variable importance ranking from a fitted
tidymodelsworkflow withvip::vip() - Explain why random
vfold_cvinflates skill estimates on spatially autocorrelated data - Diagnose spatial autocorrelation in model residuals with Moran’s I (callback to Week 3)
- Replace
vfold_cv()withspatial_block_cv()fromspatialsampleand compare honest vs. inflated metrics - Re-evaluate variable importance under spatial CV and identify features that were spatial proxies vs. real signal
Part 1: Which features mattered? ![]()
The diagnostic gap
A collect_metrics() table answers how well a model predicts.
It doesn’t answer:
- Which predictors carried the most weight?
- Would removing one of them hurt the model?
- Are the predictors the model relies on the ones that make scientific sense?
These are interpretability questions, and they’re essential before you ship a model to a stakeholder.
Note
“A model that scores well but relies on the wrong features will fail the moment conditions change.” — every applied ML scientist, eventually.
Variable Importance Plots (VIP)
A Variable Importance Plot ranks predictors by their contribution to a fitted model.
Two broad families:
- Model-specific — uses internals of the model itself
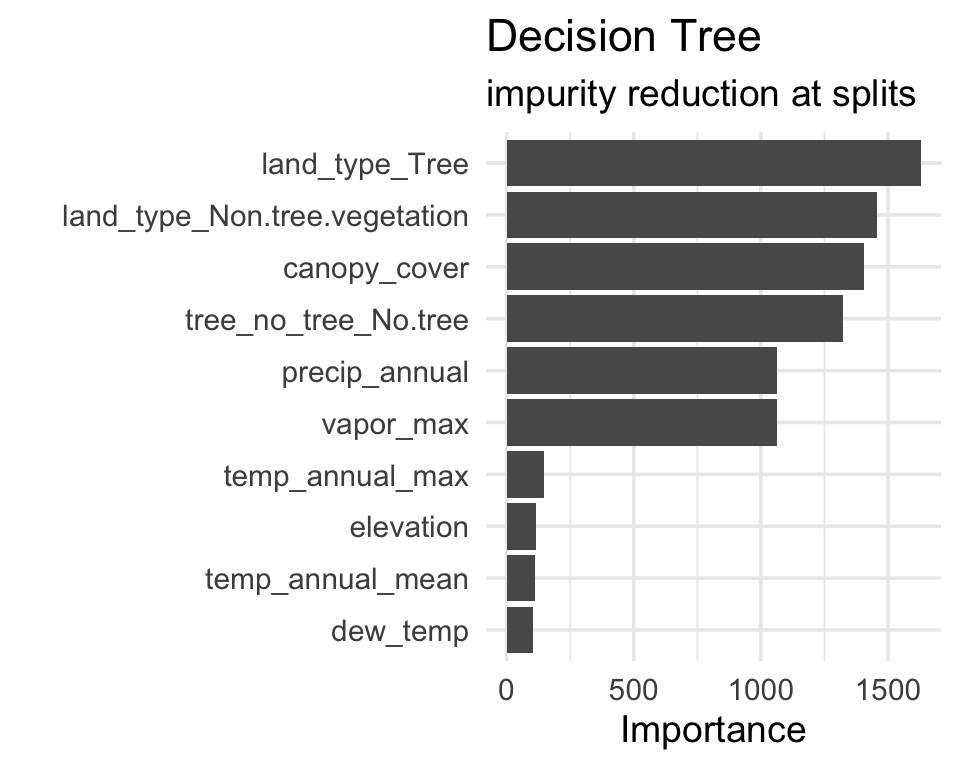
- Decision trees: the total reduction in Gini impurity at nodes that split on each variable
- Linear / logistic regression: absolute value of standardized coefficients
- Random forests / boosted trees: same as trees, averaged across the ensemble
- Model-agnostic (permutation-based) — works for any model
- Shuffle one predictor’s values, re-predict, measure how much performance drops
- If shuffling
elevationcuts AUC by 0.15,elevationis important
Important
Variable importance measures what the model used, not what is causally true. A predictor can rank high because it’s a proxy for something else (e.g., longitude encoding climate) — we’ll return to this in Part 3.
The vip package ![]()
vip is a lightweight, tidymodels-friendly interface for importance plots.
- Works with
parsnip/workflowobjects viaextract_fit_parsnip() - Supports model-specific importance for most engines
- Supports permutation importance for any model via
vip::vi_permute() vip()returns aggplot— easy to customize
Rebuild the forested pipeline ![]()
![]()
Exactly what we had last week — no new concepts yet:
set.seed(123)
forested_split <- initial_split(forested, strata = forested, prop = 0.8)
forested_train <- training(forested_split)
forested_test <- testing(forested_split)
forested_folds <- vfold_cv(forested_train, v = 10, strata = forested)
forested_recipe <- recipe(forested ~ ., data = forested_train) |>
step_impute_mean(all_numeric_predictors()) |>
step_dummy(all_nominal_predictors()) |>
step_normalize(all_numeric_predictors())Fit two models
A logistic regression (our winner from last week) and a decision tree (our runner-up):
log_mod <- logistic_reg() |>
set_engine("glm") |>
set_mode("classification")
dt_mod <- decision_tree(tree_depth = 10, min_n = 3) |>
set_engine("rpart") |>
set_mode("classification")
# Unfitted workflows (we'll reuse these for fit_resamples later)
log_wf <- workflow() |>
add_recipe(forested_recipe) |>
add_model(log_mod)
dt_wf <- workflow() |>
add_recipe(forested_recipe) |>
add_model(dt_mod)
# One-shot fit on the full training set — gives us a model we can interrogate
log_fit <- fit(log_wf, data = forested_train)
dt_fit <- fit(dt_wf, data = forested_train)Model-specific VIP ![]()
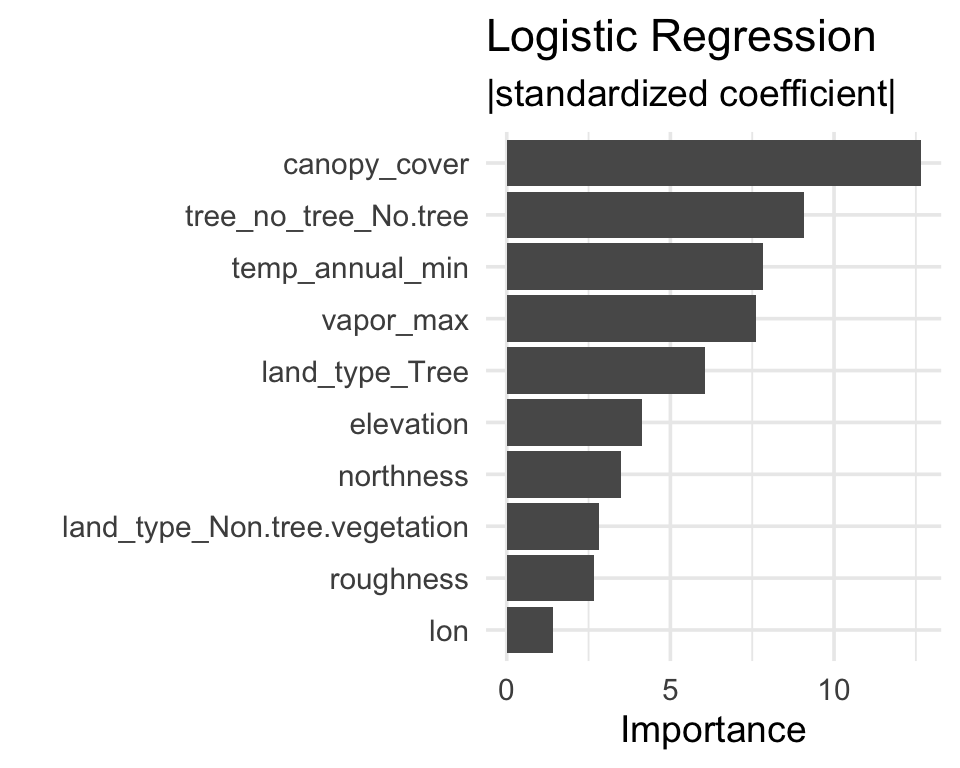
Logistic regression
Importance = absolute value of standardized coefficient.
Agreement and disagreement ![]()
A feature’s rank in each model shows where they agree and where they diverge.
# Rank each model's features,
# keep the top 10 from either
ranks <- bind_rows(
vi(extract_fit_parsnip(log_fit)) |>
mutate(model = "logistic"),
vi(extract_fit_parsnip(dt_fit)) |>
mutate(model = "tree")
) |>
group_by(model) |>
mutate(rank = rank(-Importance)) |>
group_by(Variable) |>
filter(n() == 2, min(rank) <= 10)
ggplot(ranks,
aes(model, rank, group = Variable)) +
geom_line(color = "grey70") +
geom_point(aes(color = model)) +
scale_y_reverse()- Flat lines → robust signal
- Swinging lines → model-specific
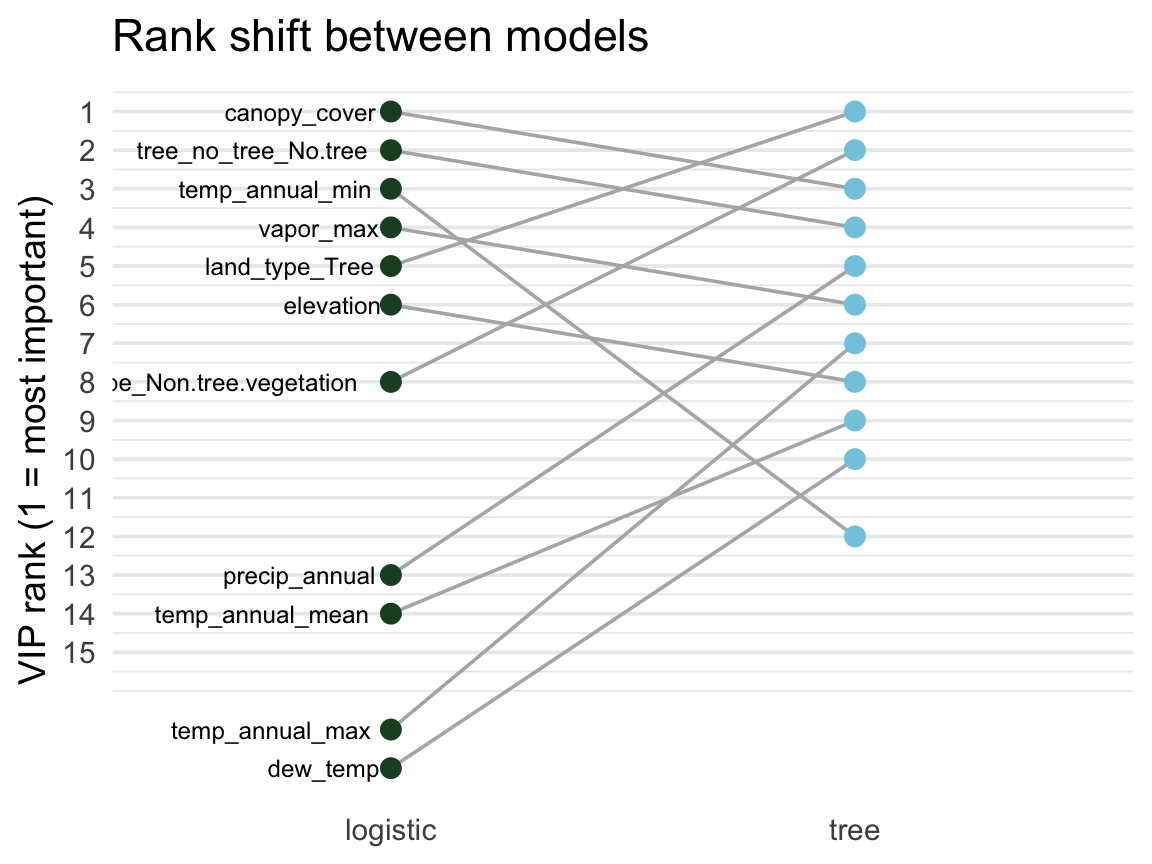
Tip
VIP is most informative across multiple models. Features important everywhere = probably real signal; features that matter only to one model type are often artifacts of how that model carves up the feature space.
Model-agnostic VIP (permutation) ![]()
Permutation importance works on any model — no internals required.
Algorithm:
- Fit the model.
- Compute a baseline metric (e.g., ROC AUC on a held-out set).
- For each predictor: randomly shuffle it, re-predict, recompute the metric.
- Importance = baseline metric − shuffled metric.
A predictor that, when scrambled, doesn’t hurt performance was not being used.
set.seed(321)
# A wrapper that returns the probability of the "Yes" class
pred_wrapper <- function(object, newdata) {
predict(object, new_data = newdata, type = "prob")$.pred_Yes
}
vip::vi_permute(
log_fit,
train = forested_train,
target = "forested",
metric = "roc_auc",
reference_class = "Yes",
pred_wrapper = pred_wrapper,
nsim = 5
) |>
vip::vip(num_features = 10) +
labs(title = "Logistic regression — permutation importance")Permutation importance is slow (shuffles + re-predicts once per feature per nsim), so you usually run it offline and save the result.
VIP takeaways
✅ VIP gives you a ranked list of what the model used
✅ Comparing model-specific VIP across several models reveals robust signals
✅ Permutation importance is model-agnostic — use it when you don’t trust a model’s internal scoring
⚠️ VIP is local to the fit you hand it:
- Fit on a different training set? Rankings can shift.
- Use a different CV scheme? Rankings can shift again.
That second point is the bridge to Part 2.
Part 2: Is my CV honest? 🗺️
What does the model actually predict? ![]()
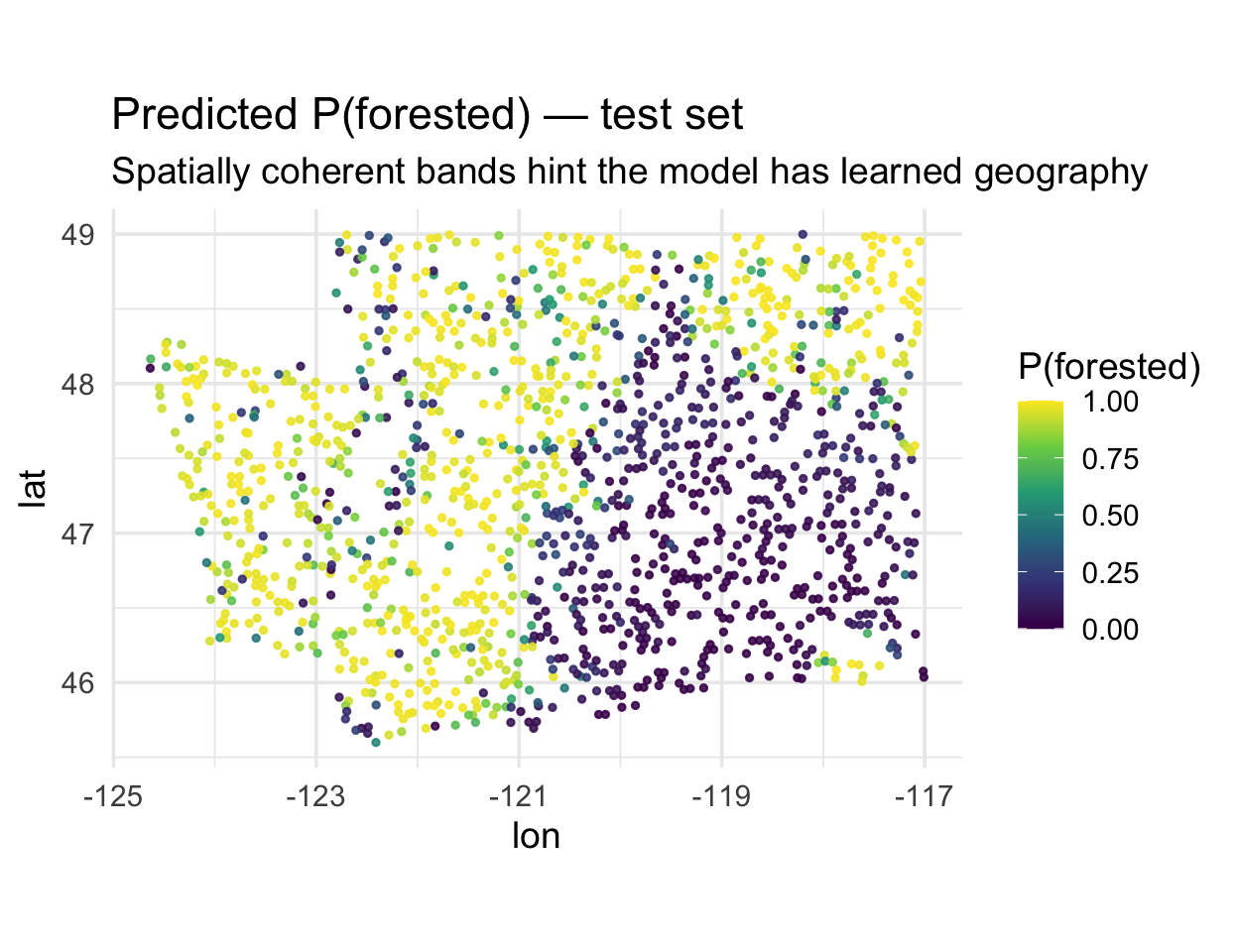
The forested dataset is spatial — map the predicted probability of forest for every test hexagon:
. . .
Notice the smooth spatial gradients. That’s informative — but also a warning sign: either (a) environmental drivers really do vary smoothly, or (b) the model is using location as a shortcut.
CV is supposed to tell us which.
Recall from Week 3: spatial autocorrelation
Are nearby values more similar than we would expect by chance?
We tested this with Moran’s I: a global summary of whether similar values cluster in space.
- positive Moran’s I → similar neighbors cluster together
- near 0 → no strong spatial pattern
- negative Moran’s I → neighbors tend to be dissimilar
And we saw that water-resources outcomes (streamflow error, nutrient concentration, flood exposure) almost always carry strong positive spatial autocorrelation.
Why this matters for ML
vfold_cv() assigns rows to folds randomly.
If a model relies on geographic interpolation, random fold assignment gives it an easy ride:
- almost every test hexagon has a training hexagon as its near neighbor
- the model can lean on local geography to predict and get rewarded at assessment time
- reported metrics reflect memorization of neighborhoods, not generalization to new places
This is a form of data leakage — but note the “if”. Whether it happens depends on your model and features: a model that has strong environmental predictors may not lean on geography at all.
Warning
The problem is hard to notice because the code still runs and the metrics are still high. You only catch it by comparing random CV to spatial CV. If they agree, you’re fine. If spatial is lower, you’ve found leakage.
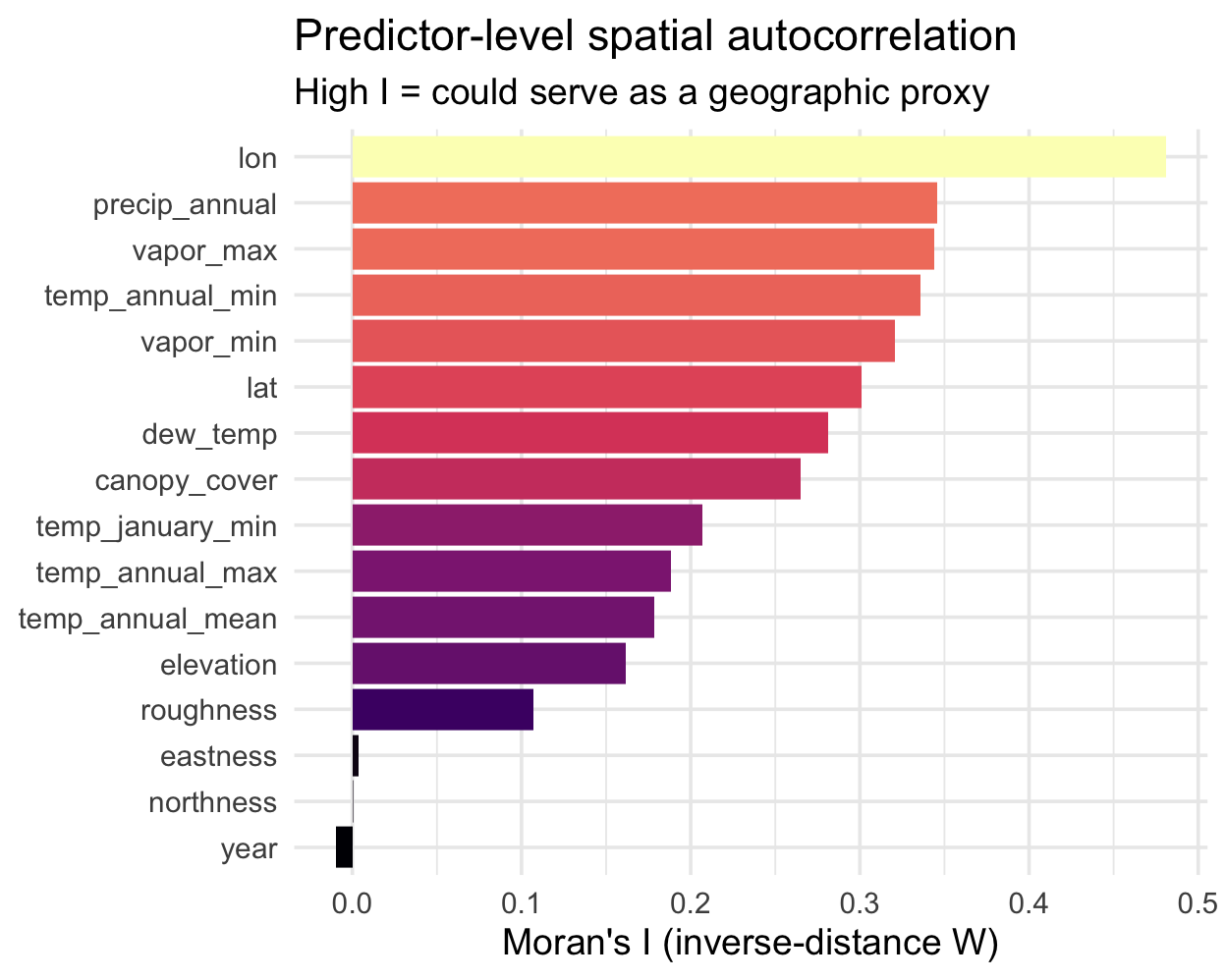
Which predictors carry geography? ![]()
Before looking at residuals, let’s use Moran’s I on the predictors themselves. Features with high spatial autocorrelation are the ones a model could use as a shortcut for location:
library(ape)
set.seed(10)
samp <- slice_sample(forested_train, n = 400)
dmat <- as.matrix(dist(cbind(samp$lon, samp$lat)))
W <- 1 / dmat; diag(W) <- 0
W <- W / rowSums(W)
preds_I <- samp |>
select(where(is.numeric), -forested) |>
map_dfr(
~ tibble(I = Moran.I(.x, W)$observed),
.id = "variable"
) |>
arrange(desc(I))Reading the ranking:
lon/lattop the list — by constructionprecip,vapor_*,temp_*are highly spatial (climate varies smoothly)eastness/northnessnear zero — aspect flips within a hillsideyear≈ 0 — it’s temporal, not spatial
Tip
Cross-reference this with your VIP plot. If your top-ranked features also sit near the top of this chart, they could be doing double duty as location encoders — a candidate for the spatial-CV check next.
Diagnosing the problem: Moran’s I on residuals
Moran’s I in Week 3 tested whether outcome values cluster.
We can also ask: do a model’s errors cluster?
If residuals are spatially autocorrelated, the model is systematically right in some areas and systematically wrong in others — it has not captured the spatial structure, it has bypassed it.
Plan:
- Attach predictions to the held-out test set
- Compute the residual (1 for a miss, 0 for a hit — this is a classification problem)
- Map the residuals
- Test for clustering with Moran’s I
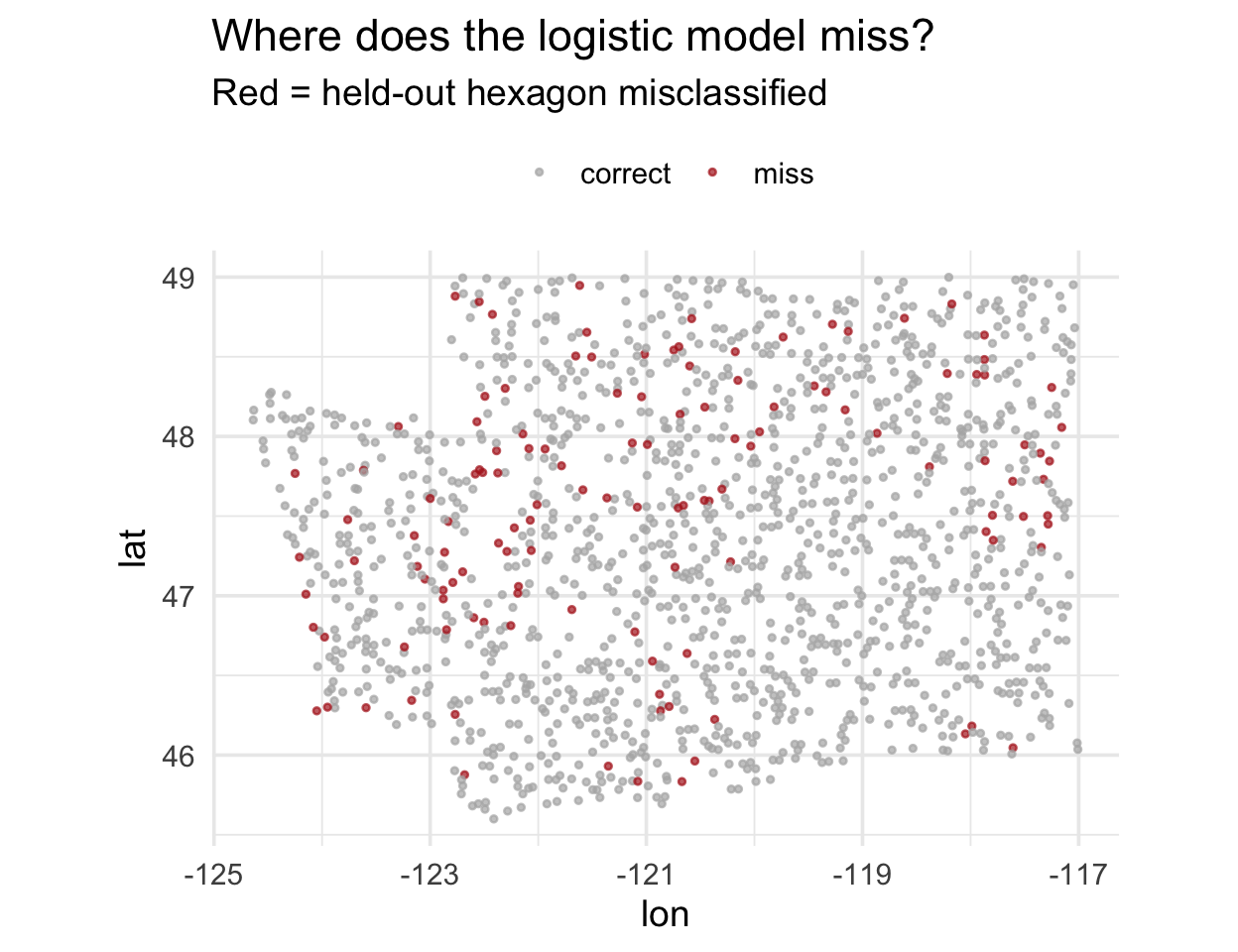
Residuals, mapped
. . .
Do you see clumps of red? That’s spatial autocorrelation in the residuals.
Quantifying it: Moran’s I on residuals
library(ape)
# Use a simple inverse-distance W on a sample of test points
set.seed(10)
samp <- preds |> slice_sample(n = 400)
dmat <- as.matrix(dist(cbind(samp$lon, samp$lat)))
W <- 1 / dmat
diag(W) <- 0
W <- W / rowSums(W)
moran_res <- Moran.I(samp$residual, W)
moran_res
#> $observed
#> [1] 0.006698239
#>
#> $expected
#> [1] -0.002506266
#>
#> $sd
#> [1] 0.004710679
#>
#> $p.value
#> [1] 0.05070528A positive, significant Moran’s I on residuals is the smoking gun: the model’s errors cluster. A small Moran’s I means the opposite — your model’s errors are well-scattered and the predictors are doing their job. Our logistic model here lands near zero, which is good news, not proof of nothing-to-see. We’ll still run spatial CV to confirm, and later in Part 4 we’ll construct a counter-example where the leakage is real.
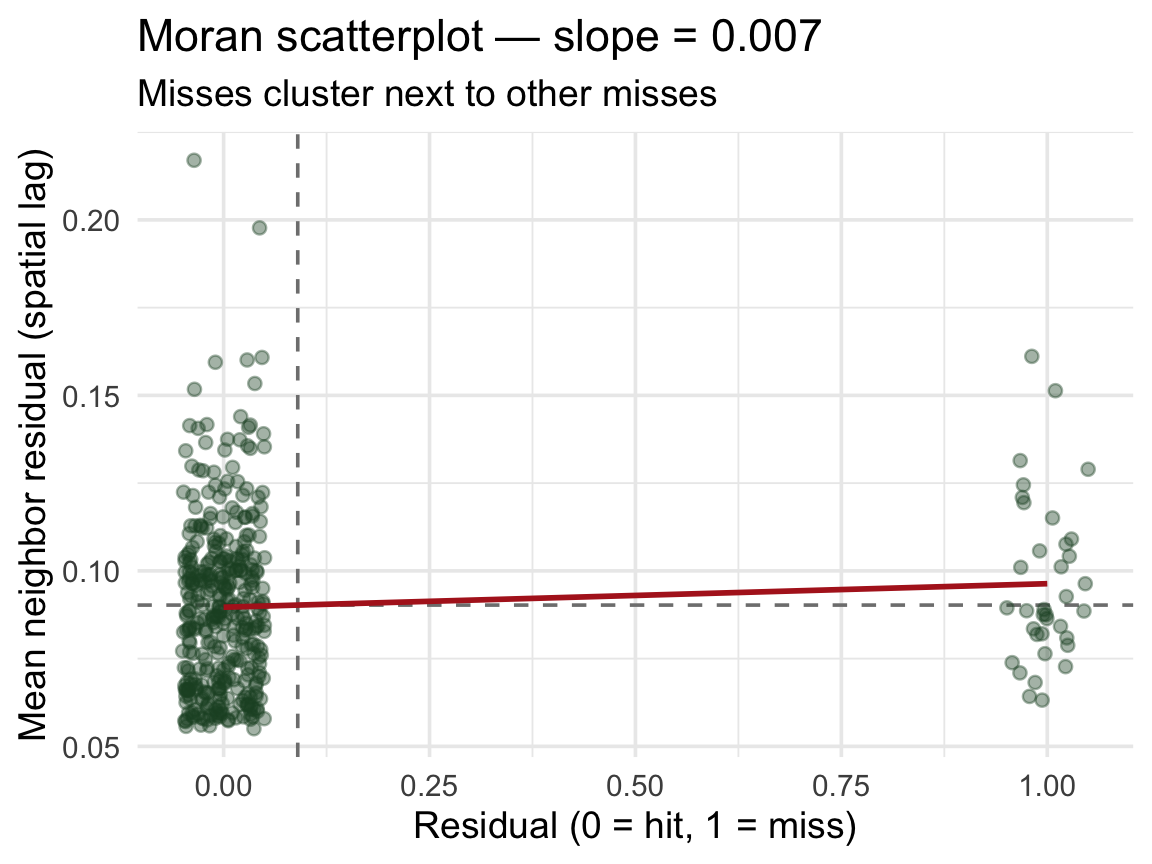
The Moran scatterplot ![]()
Moran’s I is the slope of a line through this scatterplot:
- x-axis: residual at a point
- y-axis: mean residual of its neighbors (spatial lag)
- upper-right / lower-left points → clustering
- diffuse cloud around zero → no spatial structure
. . .
The red line’s slope is Moran’s I. Positive slope → the model’s errors are geographically contagious → random CV is overstating performance.
The leakage visualized
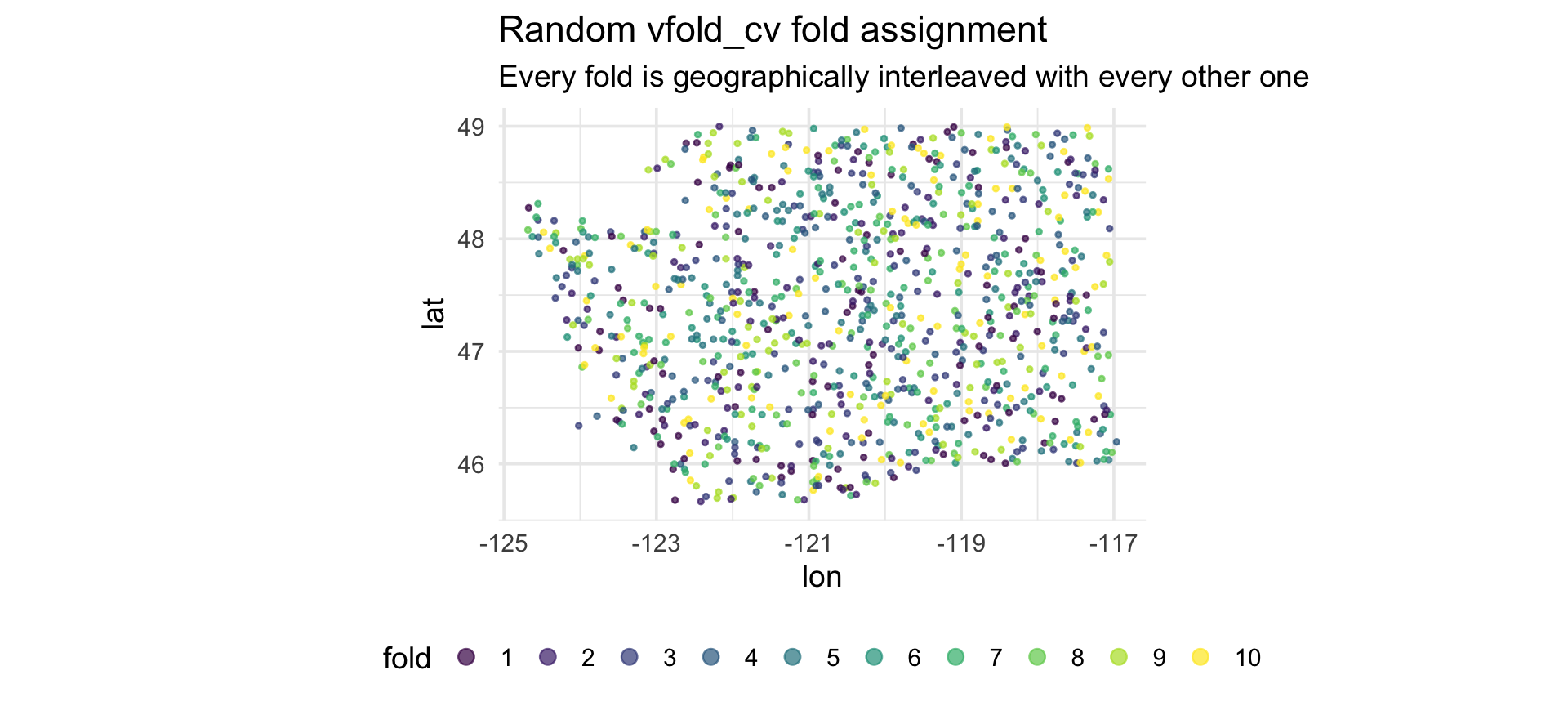
Every test point has a training point as a near neighbor. That’s the leakage.
Part 3: Spatial cross-validation ![]()
Enter spatialsample ![]()
The spatialsample package (a tidymodels extension) provides resampling schemes that respect geography.
| Function | How it splits |
|---|---|
spatial_block_cv() |
Tiles the area with a grid; assigns whole blocks to folds |
spatial_clustering_cv() |
k-means on coordinates, uses clusters as folds |
spatial_buffer_vfold_cv() |
Standard v-fold + removes a buffer of points around the held-out fold from training |
spatial_leave_location_out_cv() |
Leave out one named location (watershed, region, …) |
All produce familiar rset objects — they slot into fit_resamples() and workflow_map() unchanged.
📖 Reference: https://spatialsample.tidymodels.org/
Convert to an sf object ![]()
spatialsample needs spatial coordinates — we pass an sf object:
Note
We keep lon/lat as columns (remove = FALSE) so the model can still use them if it wants to — but conceptually that’s a smell, see Part 4.
Applying this to your data ![]()
Most real datasets don’t arrive as sf objects — they’re plain data frames with coordinate columns. Three things to watch for:
1. Column names vary. The forested data uses lon/lat, but your data might use longitude/latitude, gauge_lon/gauge_lat, X/Y, easting/northing. Pass whatever you have to coords = c(x, y):
2. The CRS must match the coordinates. Lon/lat in decimal degrees is EPSG:4326. If your data is projected (UTM, Albers, State Plane), use that EPSG code instead — otherwise your spatial blocks will be nonsense.
3. remove = FALSE is a modeling choice. Keeping the coordinate columns means your model can use them as predictors. Drop them in the recipe (step_rm(lon, lat)) if you want to force the model to rely on environmental features only.
Tip
Rule of thumb: always plot(st_geometry(your_sf)) once after conversion — if the points don’t land where you expect, the column order or CRS is wrong.
Spatial block CV ![]()
library(spatialsample)
set.seed(123)
forested_block <- spatial_block_cv(forested_sf, v = 10)
# Peek at the first few splits
print(forested_block, n = 3)
#> # 10-fold spatial block cross-validation
#> # A tibble: 10 × 2
#> splits id
#> <list> <chr>
#> 1 <split [5125/560]> Fold01
#> 2 <split [5114/571]> Fold02
#> 3 <split [5116/569]> Fold03
#> # ℹ 7 more rows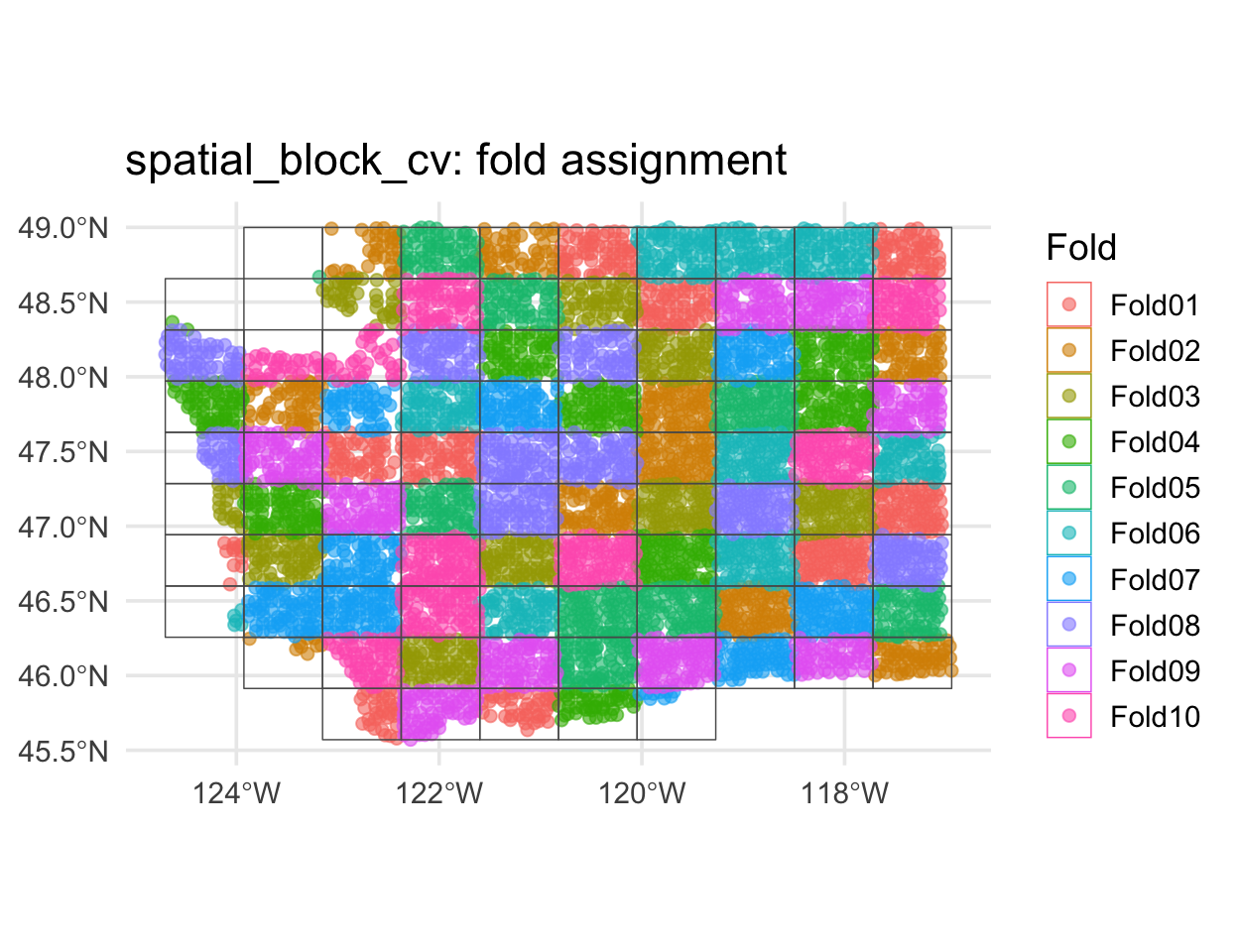
Compare the fold maps
Random vfold_cv
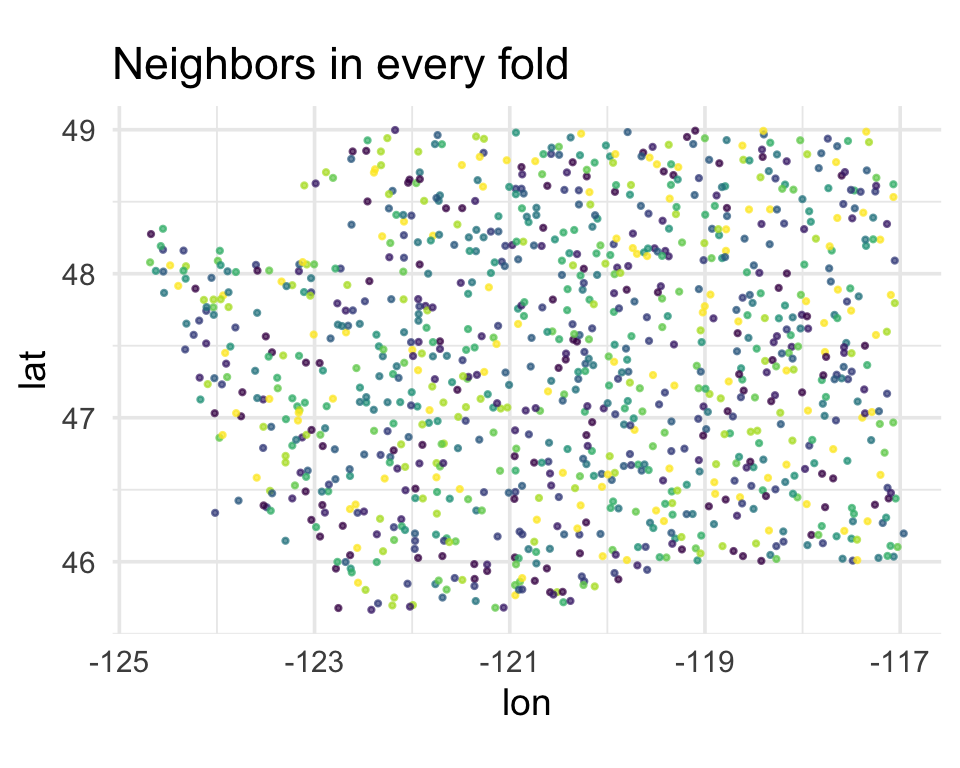
Every fold contains points from every region of Washington. A held-out point has a training point next door.
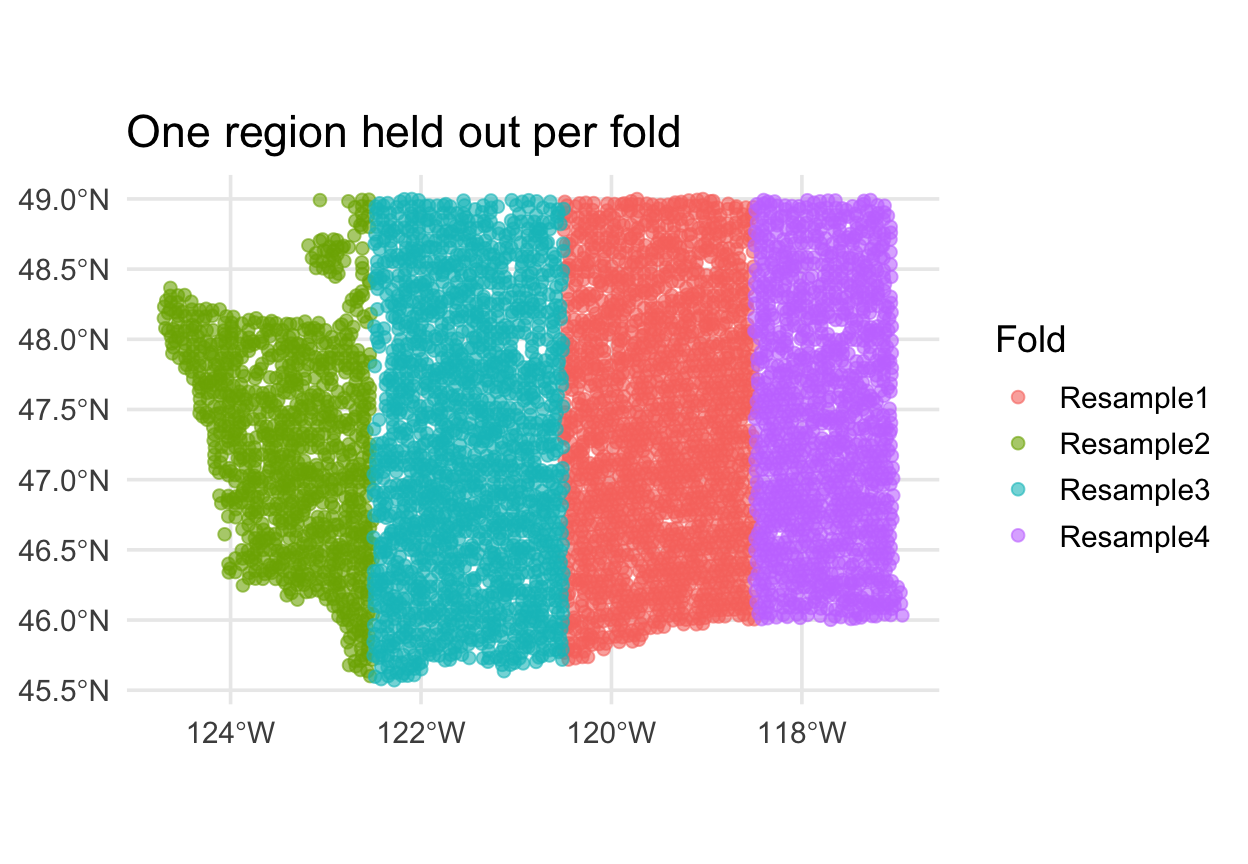
spatial_block_cv
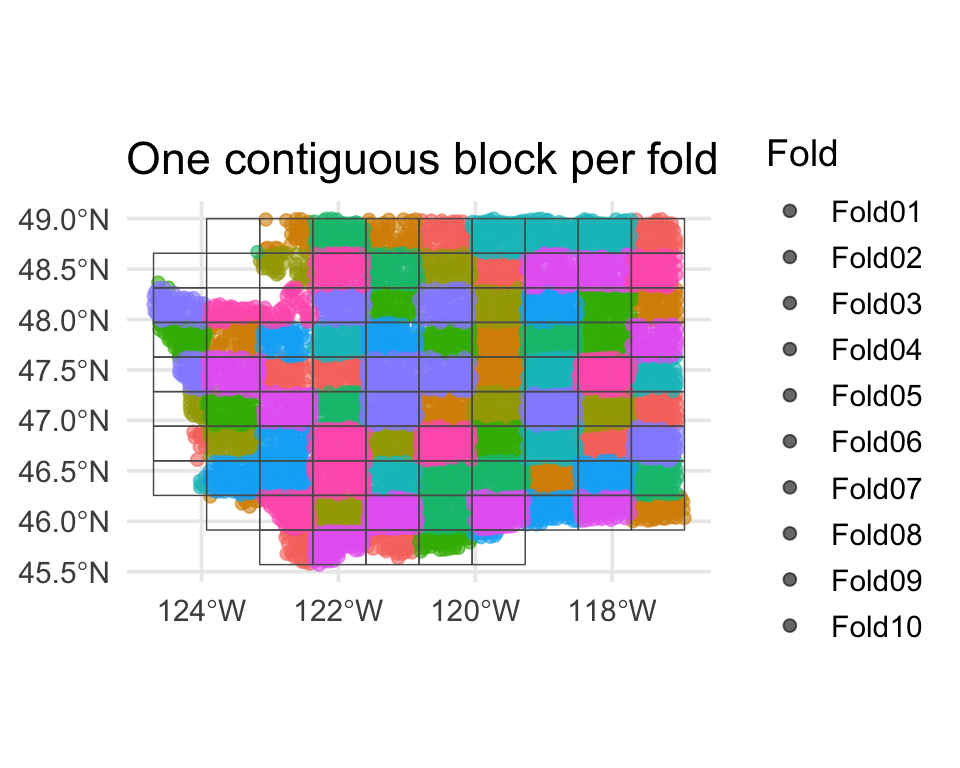

Each fold is a contiguous region. Holding it out simulates predicting somewhere the model has never seen.
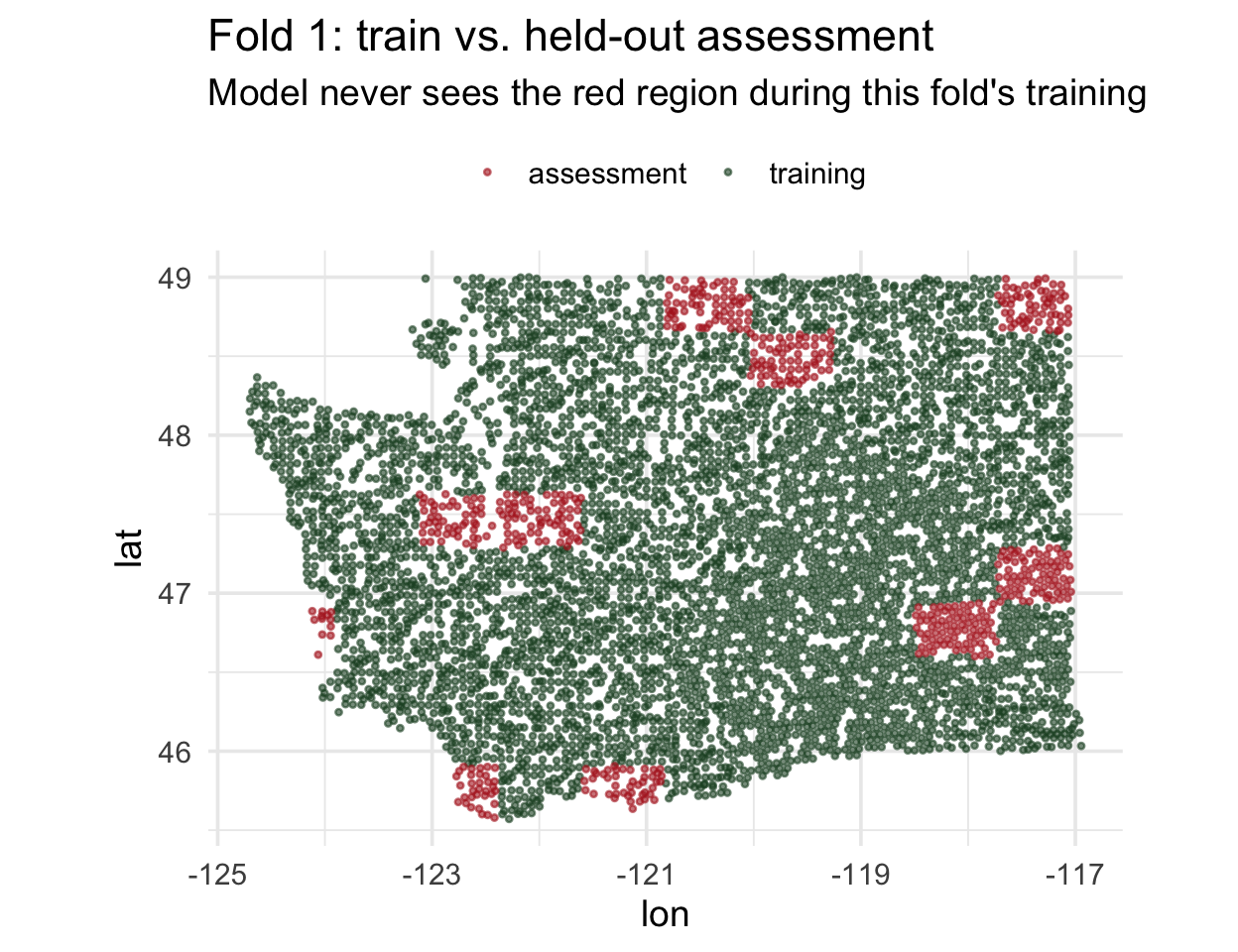
Zoom in on one fold ![]()
What does one spatial fold actually look like? Pull out fold 1 and plot its training and assessment sets:
. . .
This is the geometry random vfold_cv was quietly destroying: under random CV, those red points would have been surrounded by green ones.
Block size matters ![]()
spatial_block_cv() takes a cellsize argument (in CRS units). It controls the tile side length — and therefore how much spatial separation you actually get.
Small blocks (~0.1°)

Too small — folds interleave again, essentially random CV.
Default (~0.5°)
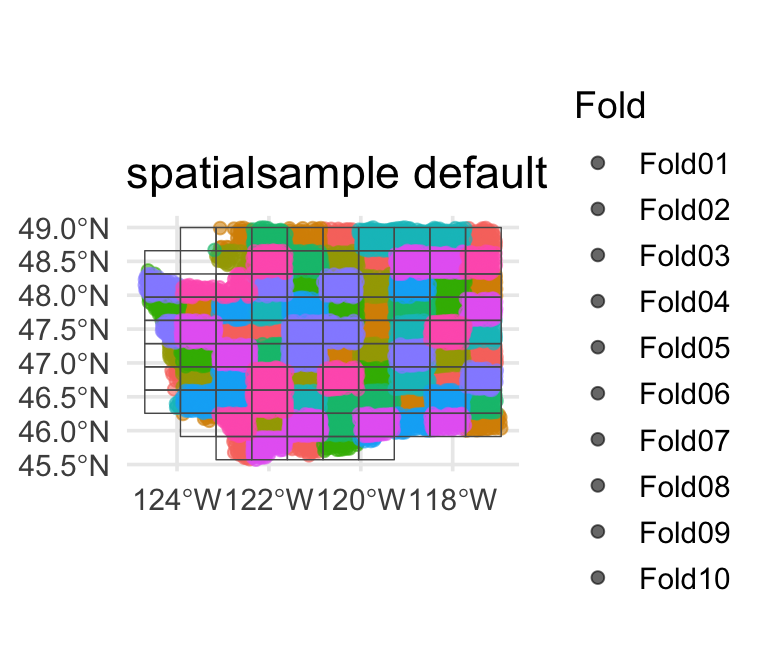
Usually a reasonable starting point.
Large blocks (~1.5°)
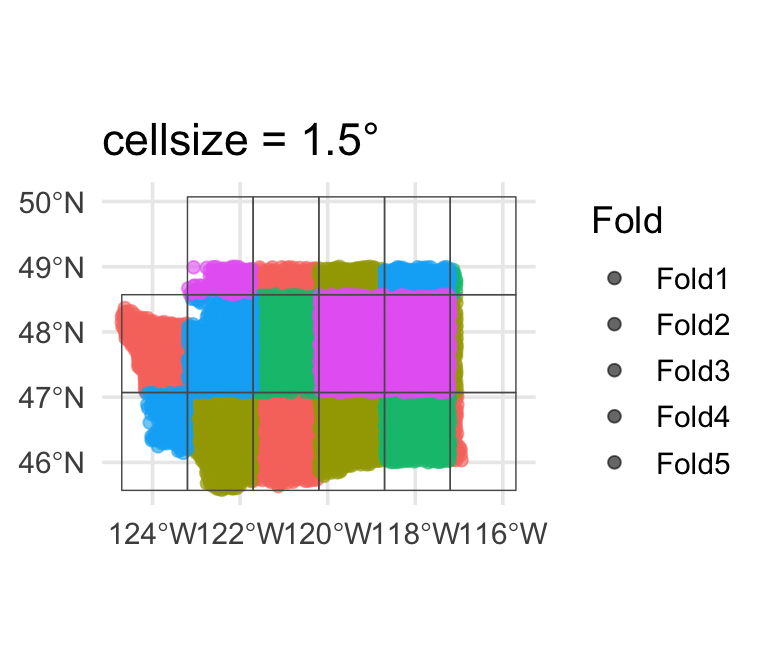
Very conservative — few effective folds, high fold-to-fold variance.
Tip
Rule of thumb: set cellsize to the spatial range of autocorrelation (from the variogram or Moran’s I distance bins). You want neighboring training points far enough from the test block that the model can’t just interpolate.
Re-fit the forested workflow ![]()
![]()
Identical to last week — only the resamples argument changes:
set.seed(2024)
cls_metrics <- metric_set(roc_auc, accuracy, brier_class)
ctrl <- control_resamples(save_pred = TRUE)
log_rs_random <- log_wf |>
fit_resamples(resamples = forested_folds, metrics = cls_metrics, control = ctrl)
log_rs_spatial <- log_wf |>
fit_resamples(resamples = forested_block, metrics = cls_metrics, control = ctrl)The moment of truth
bind_rows(
collect_metrics(log_rs_random) |> mutate(scheme = "vfold_cv (random)"),
collect_metrics(log_rs_spatial) |> mutate(scheme = "spatial_block_cv")
) |>
select(scheme, .metric, mean, std_err) |>
arrange(.metric, scheme)
#> # A tibble: 6 × 4
#> scheme .metric mean std_err
#> <chr> <chr> <dbl> <dbl>
#> 1 spatial_block_cv accuracy 0.899 0.00484
#> 2 vfold_cv (random) accuracy 0.903 0.00288
#> 3 spatial_block_cv brier_class 0.0765 0.00388
#> 4 vfold_cv (random) brier_class 0.0741 0.00148
#> 5 spatial_block_cv roc_auc 0.956 0.00365
#> 6 vfold_cv (random) roc_auc 0.959 0.00193Important
The gap is tiny — about 0.005 in ROC AUC. That’s the honest answer: on this data with these predictors, random CV was not lying. The environmental features (canopy, vapor, elevation, …) carry real transferable signal, so the model doesn’t need to memorize geography.
So does spatial CV ever matter? Yes — but only when a model is actually leaning on geography. Let’s build that model deliberately on the next slide.
What a leaky model looks like ![]()
To make spatial leakage actually happen, we’d need a model that can only use geography. Fit a decision tree on lon + lat alone — no environmental predictors:
geo_recipe <- recipe(forested ~ lon + lat, data = forested_train) |>
step_normalize(all_numeric_predictors())
geo_model <- decision_tree(tree_depth = 10, min_n = 3) |>
set_engine("rpart") |> set_mode("classification")
geo_wf <- workflow() |> add_recipe(geo_recipe) |> add_model(geo_model)
set.seed(9)
geo_rs_random <- fit_resamples(geo_wf, forested_folds, metrics = cls_metrics, control = ctrl)
geo_rs_spatial <- fit_resamples(geo_wf, forested_block, metrics = cls_metrics, control = ctrl)#> # A tibble: 4 × 4
#> scheme .metric mean std_err
#> <chr> <chr> <dbl> <dbl>
#> 1 spatial_block_cv accuracy 0.788 0.0163
#> 2 vfold_cv (random) accuracy 0.834 0.00381
#> 3 spatial_block_cv roc_auc 0.796 0.0182
#> 4 vfold_cv (random) roc_auc 0.839 0.00511Warning
Now the gap shows up. Random CV still gives the tree high marks — every held-out hexagon has a neighbor from a different fold to interpolate from. Spatial CV drops ~0.05 AUC because whole regions of Washington are held out and the model has no other signal to fall back on. That gap is spatial leakage — caught red-handed.
ROC curves: all four fits overlaid ![]()
Overlay the out-of-fold predictions from each model × CV scheme:
roc_data <- bind_rows(
collect_predictions(log_rs_random) |>
mutate(model = "all features",
scheme = "random"),
collect_predictions(log_rs_spatial) |>
mutate(model = "all features",
scheme = "spatial"),
collect_predictions(geo_rs_random) |>
mutate(model = "geo-only",
scheme = "random"),
collect_predictions(geo_rs_spatial) |>
mutate(model = "geo-only",
scheme = "spatial")
) |>
group_by(model, scheme) |>
roc_curve(truth = forested, .pred_Yes)
ggplot(roc_data,
aes(1 - specificity, sensitivity,
color = scheme, linetype = model)) +
geom_path(linewidth = 1.1) +
coord_equal()The all-features pair is on top of each other — no leakage to find. The geo-only pair shows visible daylight between random and spatial — the spatial curve sagging is what leakage looks like.
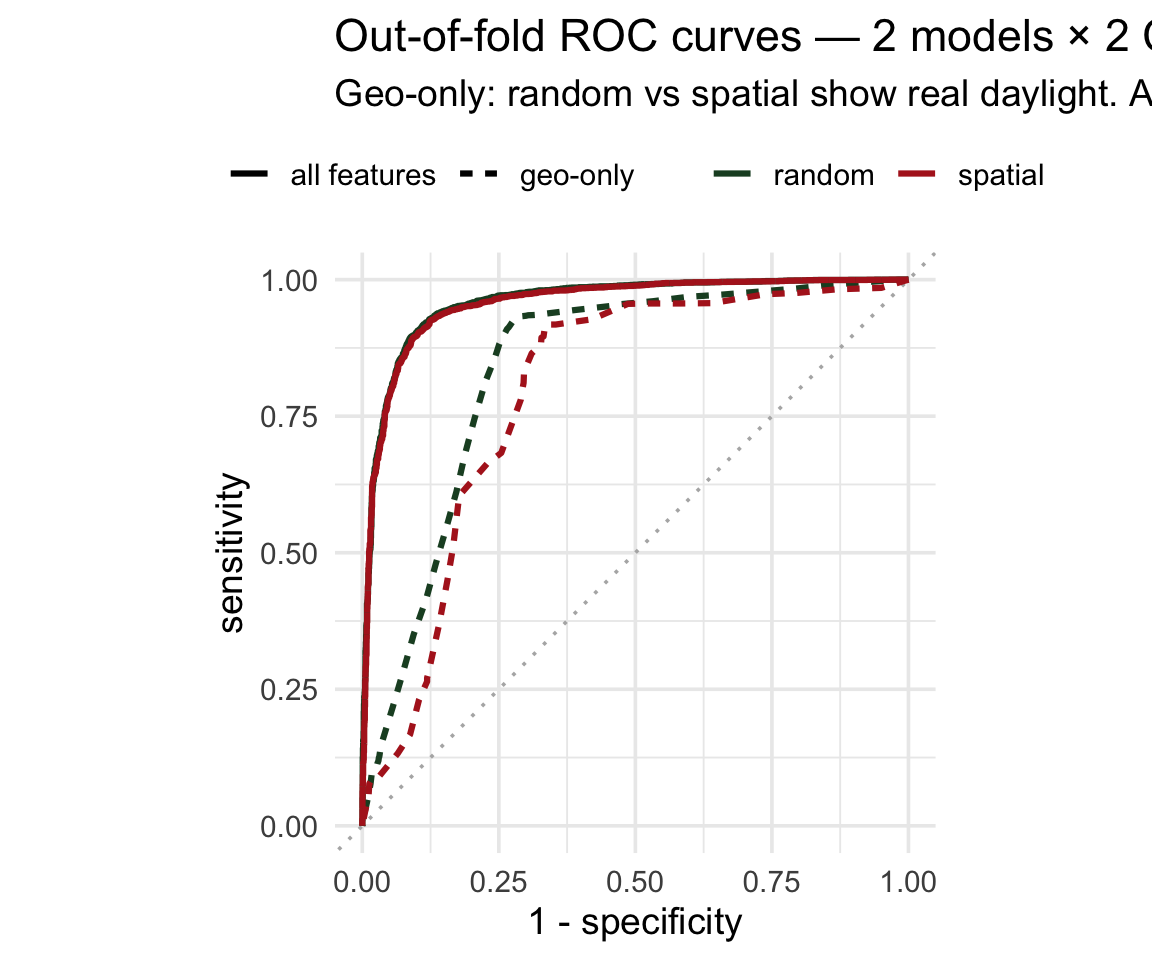
It’s not just lower — it’s more variable ![]()
The mean hides the story. Pull per-fold metrics for both models × both schemes:
per_fold <- bind_rows(
collect_metrics(log_rs_random, summarize = FALSE) |>
mutate(model = "all features", scheme = "random"),
collect_metrics(log_rs_spatial, summarize = FALSE) |>
mutate(model = "all features", scheme = "spatial"),
collect_metrics(geo_rs_random, summarize = FALSE) |>
mutate(model = "geo-only", scheme = "random"),
collect_metrics(geo_rs_spatial, summarize = FALSE) |>
mutate(model = "geo-only", scheme = "spatial")
) |> filter(.metric == "roc_auc")
ggplot(per_fold,
aes(interaction(model, scheme), .estimate,
color = scheme)) +
geom_boxplot() +
geom_jitter(width = 0.08)All-features: both CV schemes give tight, near-identical distributions → no leakage. Geo-only: spatial folds show lower mean AND wider spread — the model’s performance depends heavily on which region you hold out.
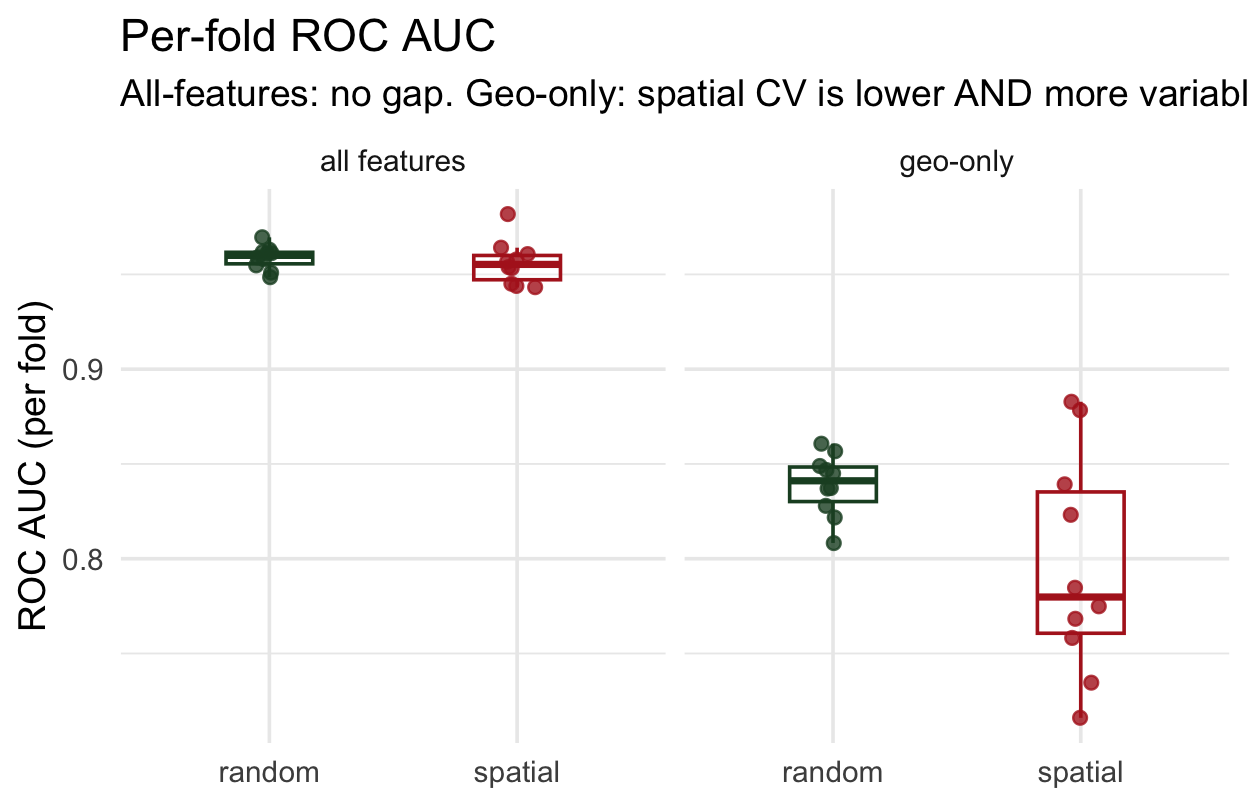
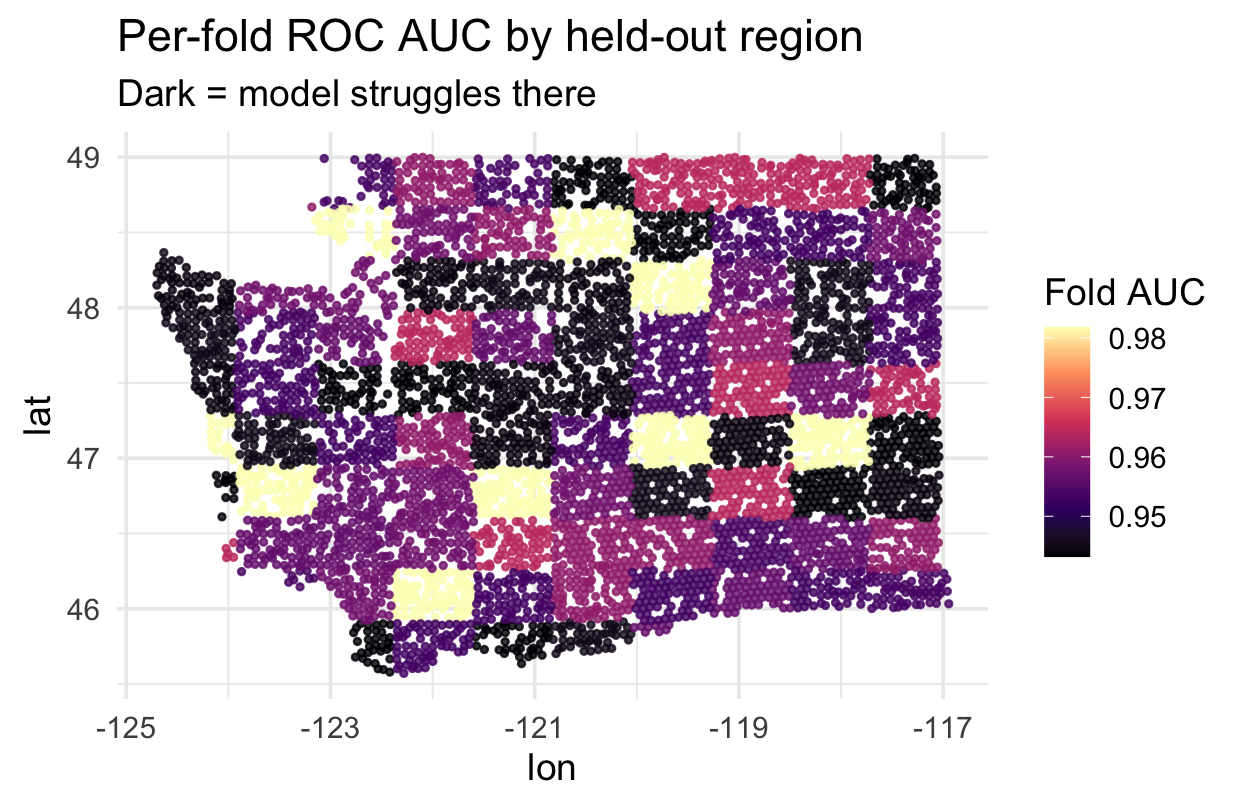
Where does spatial CV struggle? ![]()
Each spatial fold’s held-out region has a location. Map each fold’s AUC to see which regions are hardest to predict:
fold_auc <- collect_metrics(
log_rs_spatial, summarize = FALSE) |>
filter(.metric == "roc_auc") |>
select(id, fold_auc = .estimate)
# join each fold's AUC back to its
# held-out coordinates (see repo)
ggplot(fold_locations,
aes(lon, lat, color = fold_auc)) +
geom_point() +
scale_color_viridis_c(option = "magma")Regions where AUC is lowest are where your model has the least transferable signal — a scientific finding telling you where to invest more training data or features.
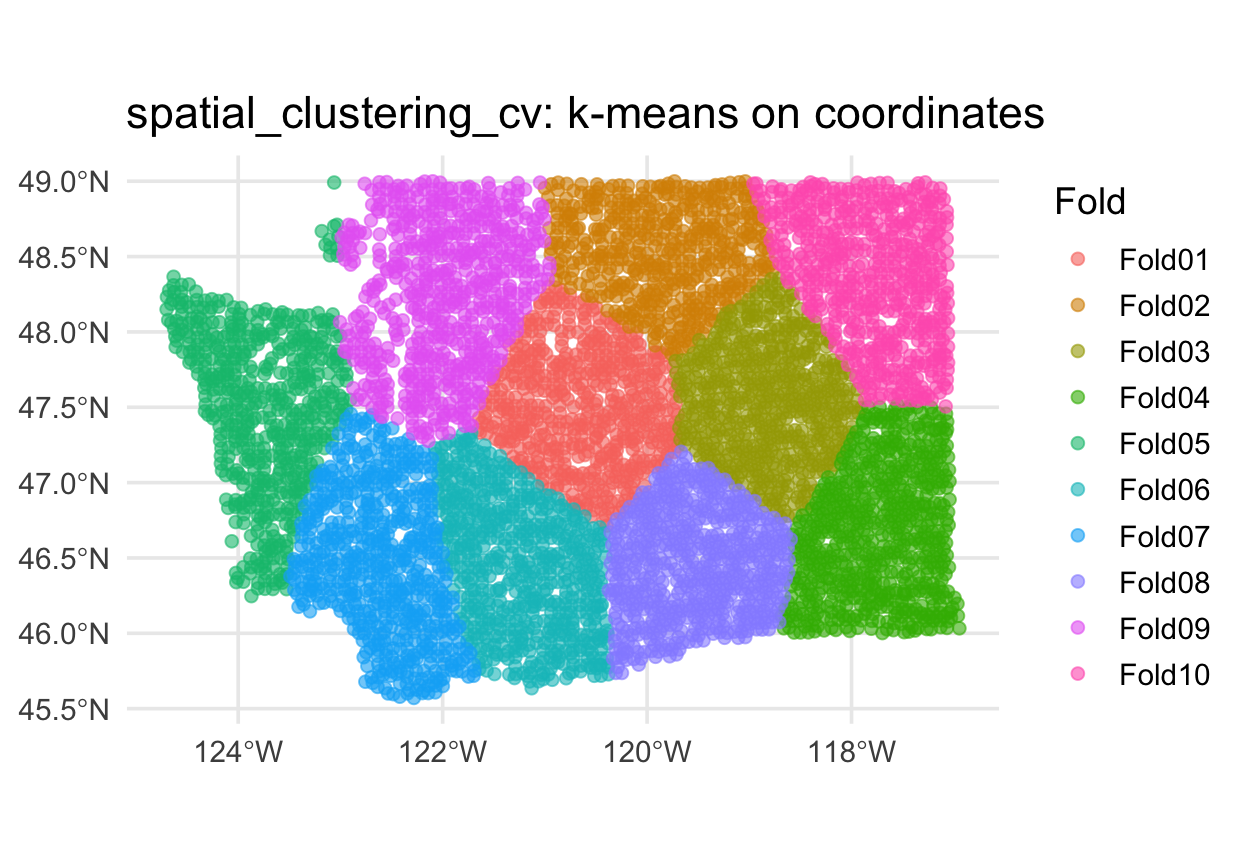
Alternative: spatial_clustering_cv ![]()
When blocks don’t fit your data (e.g., irregularly shaped study regions), cluster-based folds are often better:
Alternative: buffered v-fold ![]()
What if you want random folds but guarantee no training point is within, say, 10 km of a test point?
- Each training set has points within
bufferof held-out points removed - Preserves fold size randomness; enforces a minimum spatial gap
- Good compromise when blocks are too coarse but random CV is too leaky
Warning
buffer is expressed in the units of the CRS. Buffers on geographic (lon/lat) CRSs give meaningless results — always transform to a projected CRS first.
Alternative: leave-location-out CV ![]()
For water resources, the natural unit is usually a watershed, gauge basin, or ecoregion — not a square block. spatial_leave_location_out_cv() holds out one named group at a time:
Each fold trains on 3 regions and predicts the 4th — answers “could I predict eastern WA from coastal WA?”, which vfold_cv can’t. In your project, swap region for HUC8, gauge basin, or climate zone.
When not to reach for spatial CV 🤔
Spatial CV is not a universal “do this instead of vfold_cv” replacement. It depends on your scientific question:
✅ Use spatial CV when:
- You want to generalize to new locations (operational forecasting, transfer to a new study region, national-scale extrapolation)
- Your test set will be geographically distinct from training data
- You have significant spatial autocorrelation in residuals (check it!)
⚠️ Random CV can still be defensible when:
- Your question is explicitly “predict well in the same places and periods I have training data” (e.g., gap-filling a sensor network where every test point has dense neighbors by design)
- Your data is not spatially structured (rare in water resources)
- You want a pre-deployment upper-bound estimate and are planning to re-evaluate on independent regions later
Tip
A mature workflow often reports both: random CV as a best-case ceiling, spatial CV as a generalization floor. The gap between them is informative — a large gap is a flag that your model is leaning hard on geography.
Part 4: Trust your diagnostics
Putting the pieces together
We now have two models under two CV schemes — that’s a 2×2 we can reason about:
- All-features model under random CV vs spatial CV — gap is tiny → predictors carry real signal
- Geo-only model under random CV vs spatial CV — gap is large → model is leaning on geography
The size of the gap between random and spatial CV is a diagnostic of how much the model depends on geographic shortcuts. You compute it, read it, and decide whether your model is trustworthy on new geography.
VIP told us what the model used. Spatial CV told us whether those features generalize. Together they answer “can I trust this model outside the training area?”
The 2×2 comparison
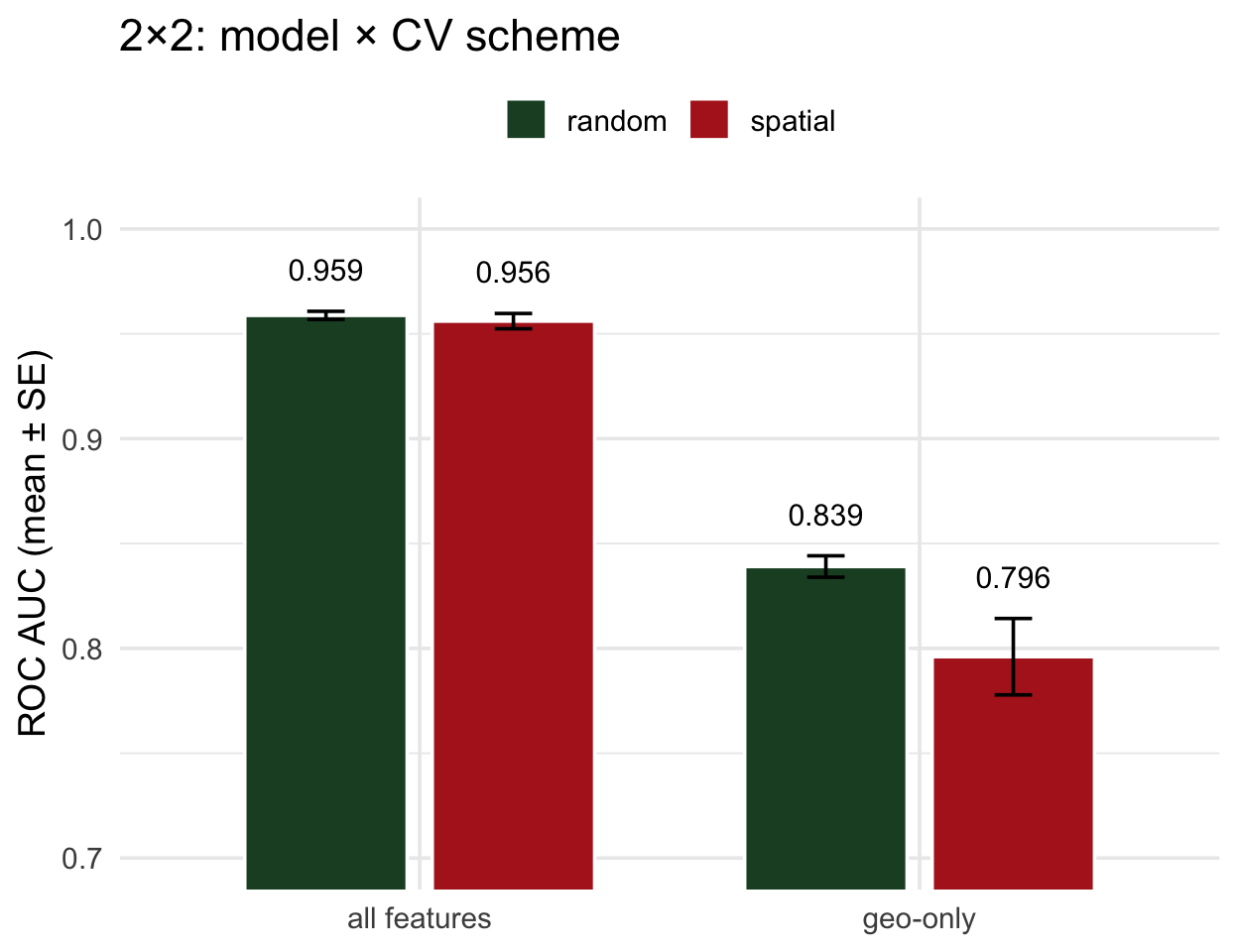
- All features, random ≈ spatial → no leakage; random CV was telling the truth
- Geo-only, random ≫ spatial → the gap is the leakage
- The diagnostic is the within-model gap, not the between-model one
. . .
Read it this way: compare the two bars within each model. Equal = trust the number. Unequal = your model’s random-CV score is inflated by exactly that much.
The same idea in regression ![]()
Every example above used classification (forested: Yes/No) — but the same leakage story applies to regression. You just swap the metric.
Pretend we’re predicting log mean streamflow (logQmean) from basin attributes for CAMELS gauges — a spatial regression problem. The 2×2 would be built the same way:
- Classification: higher ROC AUC = better → spatial CV usually lowers AUC
- Regression: higher R² = better → spatial CV usually lowers R²
- Regression: lower RMSE = better → spatial CV usually raises RMSE
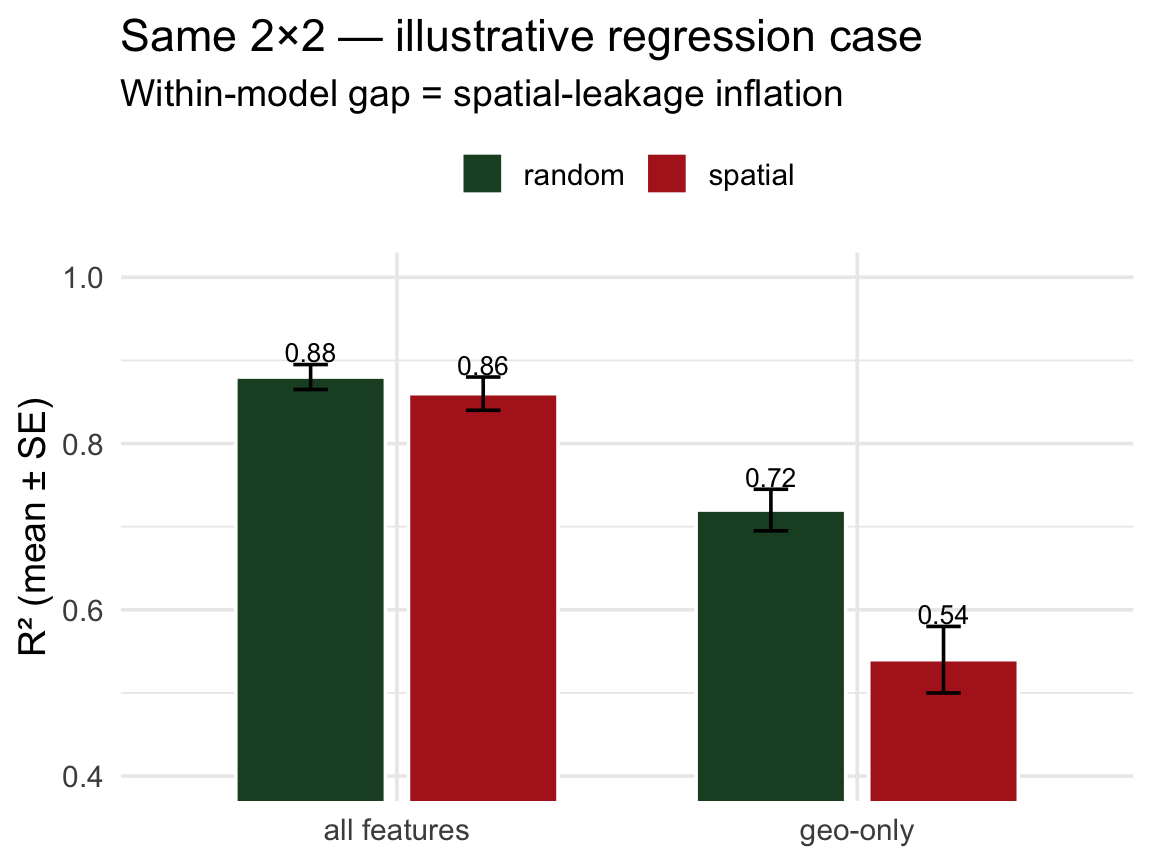
The within-model gap (random vs spatial) is the diagnostic — just on a regression metric instead of AUC.
Important
For your lab: CAMELS basins have gauge_lon/gauge_lat. Random vfold_cv on CAMELS is the regression version of the story above. Build a spatial_block_cv next to it and report both — the delta is the honest cost of generalization.
What this tells us
The within-model gap between random and spatial CV is a diagnostic: it measures how much the model’s performance depended on geographic interpolation.
- Small gap (our all-features model): the predictors carry transferable signal — random CV was honest.
- Large gap (our geo-only model): performance was inflated by geography — trust spatial CV.
Tip
A practical workflow:
- Fit your model. Look at VIP → identify your top features.
- Run both random and spatial CV.
- If the two CV scores agree → random CV was telling the truth, ship it.
- If spatial CV is meaningfully lower → your model is leaning on geography. Suspect the top features that rank in VIP — they may be spatial proxies (elevation as climate, etc). Consider removing them and refitting.
Takeaways
collect_metrics()is necessary, not sufficient. You also need to ask what the model uses and whether your evaluation respects the data’s structure.vipgives you the “what”. Model-specific importance for common engines, permutation importance for anything else.Spatial CV is a diagnostic, not a punishment. Sometimes the gap is tiny (our all-features model) — that’s evidence your predictors generalize. Sometimes it’s huge (our geo-only model) — that’s the leakage being caught.
spatialsamplegives you the other CV scheme.spatial_block_cv,spatial_clustering_cv, and buffered variants all slot into the existing tidymodels pipeline.The two tools must be used together. VIP alone tells you what the model used; the random-vs-spatial CV gap tells you whether those features generalize.
Looking ahead
Week 6: We’ll return to the forested problem with a full zoo of model families (random forests, boosted trees, SVMs, neural nets) — and hyperparameter tuning. Everything you learned today about honest evaluation still applies.
For your project: any time your data is spatial (which in water resources is always), default to spatial_block_cv or spatial_clustering_cv. The cost is a few extra lines; the benefit is numbers you can defend.
📖 Further reading:
spatialsample: https://spatialsample.tidymodels.org/vip: https://koalaverse.github.io/vip/- Roberts et al. (2017), Cross-validation strategies for data with temporal, spatial, hierarchical, or phylogenetic structure
- Meyer et al. (2019), Importance of spatial predictor variable selection in machine learning applications