Basic Use
Mike Johnson
Lynker, NOAA-Affiliatebasic_use.Rmd
library(hydrofabric3D)The goal of hydrofabric3D is to generate DEM-based cross sections for hydrographic networks.
Installation
You can install the development version of hydrofabric3D from GitHub with:
# install.packages("devtools")
devtools::install_github("mikejohnson51/hydrofabric3D")Example
This is a basic example which shows you how to cut cross sections for a network.
Define Network
library(hydrofabric3D)
library(dplyr)
#>
#> Attaching package: 'dplyr'
#> The following objects are masked from 'package:stats':
#>
#> filter, lag
#> The following objects are masked from 'package:base':
#>
#> intersect, setdiff, setequal, union
(net = linestring %>%
mutate(bf_width = exp(0.700 + 0.365* log(totdasqkm))))
#> Simple feature collection with 325 features and 5 fields
#> Geometry type: LINESTRING
#> Dimension: XY
#> Bounding box: xmin: 77487.09 ymin: 890726.5 xmax: 130307.4 ymax: 939129.8
#> Projected CRS: NAD83 / Conus Albers
#> # A tibble: 325 × 6
#> nhdplus_comid geometry comid totdasqkm dist_m bf_width
#> * <chr> <LINESTRING [m]> <dbl> <dbl> <dbl> <dbl>
#> 1 101 (128525.6 892408.3, 128565.7 … 1.01e2 7254. 3.25e3 51.7
#> 2 24599575 (128084.7 892952.4, 128525.6 … 2.46e7 7249. 7.00e2 51.6
#> 3 1078635 (127687.6 893270.4, 127799.7 … 1.08e6 7248. 5.22e2 51.6
#> 4 1078637 (124942.8 893959.6, 124948.2 … 1.08e6 68.2 4.17e3 9.41
#> 5 1078639 (125523.1 892528, 125657.3 89… 1.08e6 7180. 2.76e3 51.5
#> 6 1078577 (123219.9 902292.8, 123233.5 … 1.08e6 19.8 9.91e3 5.99
#> 7 1078575 (121975.5 909050.8, 122028.9 … 1.08e6 41.3 1.87e4 7.83
#> 8 1078657 (124263.8 892410.4, 124420.6 … 1.08e6 7179. 1.66e3 51.5
#> 9 1078663 (125628.9 892216, 125555.7 89… 1.08e6 0.099 7.54e2 0.866
#> 10 1078643 (124248.1 892440.7, 124263.8 … 1.08e6 7178. 3.41e1 51.5
#> # ℹ 315 more rows
plot(net$geometry)
Cut cross sections
(transects = cut_cross_sections(net = net,
id = "comid",
cs_widths = pmax(50, net$bf_width * 7),
num = 10) )
#> Smoothing
#> Densifying
#> Cutting
#> Formating
#> Warning: st_centroid assumes attributes are constant over geometries
#> Simple feature collection with 2275 features and 7 fields
#> Geometry type: LINESTRING
#> Dimension: XY
#> Bounding box: xmin: 77473.82 ymin: 890553.2 xmax: 130336.7 ymax: 939136.7
#> Projected CRS: NAD83 / Conus Albers
#> # A tibble: 2,275 × 8
#> ds_distance cs_measure hy_id cs_widths geometry cs_id
#> <dbl> <dbl> <dbl> <dbl> <LINESTRING [m]> <int>
#> 1 367. 11.2 101 362. (128409.2 892046.4, 128768.1… 1
#> 2 818. 25.0 101 362. (128553.4 891572.7, 128890.1… 2
#> 3 1242. 38.0 101 362. (128838.4 891220.8, 129145.3… 3
#> 4 1629. 49.8 101 362. (129198.3 890891.1, 129319.4… 4
#> 5 1962. 60.0 101 362. (129590.7 891044.8, 129463.9… 5
#> 6 2241. 68.5 101 362. (129732.4 891134.4, 129742 8… 6
#> 7 2487. 76.0 101 362. (129719.7 891083.8, 130016.4… 7
#> 8 2813. 86.0 101 362. (129907.8 891136.4, 130208.7… 8
#> 9 3260. 99.6 101 362. (130254.3 890553.2, 130336.7… 9
#> 10 77.8 11.1 24599575 362. (127993.3 892778.2, 128274.1… 1
#> # ℹ 2,265 more rows
#> # ℹ 2 more variables: lengthm <dbl>, sinuosity <dbl>
plot(transects$geometry)
Define Cross section points
(pts = cross_section_pts(transects,
dem = "/vsicurl/https://prd-tnm.s3.amazonaws.com/StagedProducts/Elevation/1/TIFF/USGS_Seamless_DEM_1.vrt"))
#> Simple feature collection with 23416 features and 11 fields
#> Geometry type: POINT
#> Dimension: XY
#> Bounding box: xmin: 77475.72 ymin: 890567.9 xmax: 130333.3 ymax: 939134.7
#> Projected CRS: NAD83 / Conus Albers
#> # A tibble: 23,416 × 12
#> hy_id cs_id pt_id Z lengthm relative_distance ds_distance cs_measure
#> <dbl> <int> <int> <dbl> <dbl> <dbl> <dbl> <dbl>
#> 1 101 1 1 42.2 362. 0 367. 11.2
#> 2 101 1 2 42.1 362. 32.9 367. 11.2
#> 3 101 1 3 42.5 362. 65.7 367. 11.2
#> 4 101 1 4 42.4 362. 98.6 367. 11.2
#> 5 101 1 5 40.2 362. 131. 367. 11.2
#> 6 101 1 6 40.2 362. 164. 367. 11.2
#> 7 101 1 7 36.2 362. 197. 367. 11.2
#> 8 101 1 8 39.9 362. 230. 367. 11.2
#> 9 101 1 9 39.7 362. 263. 367. 11.2
#> 10 101 1 10 40.5 362. 296. 367. 11.2
#> # ℹ 23,406 more rows
#> # ℹ 4 more variables: cs_widths <dbl>, sinuosity <dbl>, points_per_cs <dbl>,
#> # geometry <POINT [m]>Classify Cross section points
(classified_pts = classify_points(pts))
#> Simple feature collection with 23416 features and 7 fields
#> Geometry type: POINT
#> Dimension: XY
#> Bounding box: xmin: 77475.72 ymin: 890567.9 xmax: 130333.3 ymax: 939134.7
#> Projected CRS: NAD83 / Conus Albers
#> # A tibble: 23,416 × 8
#> hy_id cs_id pt_id Z relative_distance cs_widths class
#> <dbl> <int> <int> <dbl> <dbl> <dbl> <chr>
#> 1 101 1 1 42.2 0 362. left_bank
#> 2 101 1 2 42.2 32.9 362. left_bank
#> 3 101 1 3 42.3 65.7 362. left_bank
#> 4 101 1 4 41.7 98.6 362. channel
#> 5 101 1 5 40.9 131. 362. channel
#> 6 101 1 6 38.9 164. 362. channel
#> 7 101 1 7 36.2 197. 362. bottom
#> 8 101 1 8 38.6 230. 362. channel
#> 9 101 1 9 40.0 263. 362. right_bank
#> 10 101 1 10 41.2 296. 362. right_bank
#> # ℹ 23,406 more rows
#> # ℹ 1 more variable: geometry <POINT [m]>Explore!
library(ggplot2)
ggplot(data = filter(classified_pts, hy_id == 101) ) +
geom_line(aes(x = relative_distance, y = Z)) +
geom_point(aes(x = relative_distance, y = Z, color = class)) +
facet_wrap(~cs_id, scales = "free") +
theme_minimal() +
theme(legend.position = "bottom")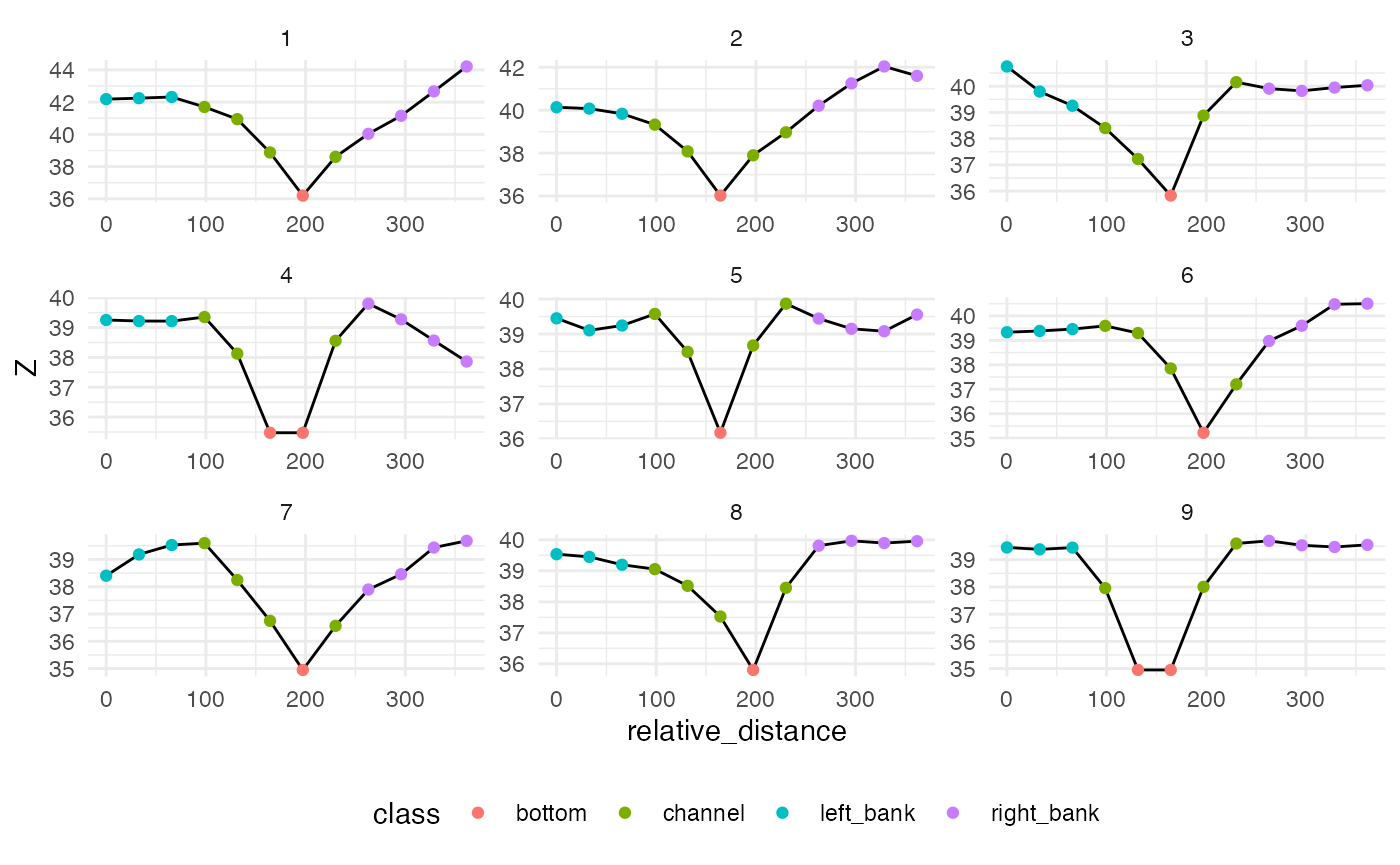
Time to get 2275 transects and 23416 classified points …
system.time({
cs = net %>%
cut_cross_sections(id = "comid",
cs_widths = pmax(50, net$bf_width * 7),
num = 10) %>%
cross_section_pts(dem = '/vsicurl/https://prd-tnm.s3.amazonaws.com/StagedProducts/Elevation/1/TIFF/USGS_Seamless_DEM_1.vrt') %>%
classify_points()
})
#> Smoothing
#> Densifying
#> Cutting
#> Formating
#> Warning: st_centroid assumes attributes are constant over geometries
#> user system elapsed
#> 18.788 0.339 21.747